Elastic Maps and Nets for Approximating Principal Manifolds and Their Application to Microarray Data Visualization
Principal manifolds are defined as lines or surfaces passing through the middle'' of data distribution. Linear principal manifolds (Principal Components Analysis) are routinely used for dimension reduction, noise filtering and data visualization. Recently, methods for constructing non-linear principal manifolds were proposed, including our elastic maps approach which is based on a physical analogy with elastic membranes. We have developed a general geometric framework for constructing principal objects’’ of various dimensions and topologies with the simplest quadratic form of the smoothness penalty which allows very effective parallel implementations. Our approach is implemented in three programming languages (C++, Java and Delphi) with two graphical user interfaces (VidaExpert http://bioinfo.curie.fr/projects/vidaexpert and ViMiDa http://bioinfo-out.curie.fr/projects/vimida applications). In this paper we overview the method of elastic maps and present in detail one of its major applications: the visualization of microarray data in bioinformatics. We show that the method of elastic maps outperforms linear PCA in terms of data approximation, representation of between-point distance structure, preservation of local point neighborhood and representing point classes in low-dimensional spaces.
💡 Research Summary
The paper introduces the Elastic Map method as a physically‑inspired framework for constructing non‑linear principal manifolds—geometric objects that pass through the “middle” of a data distribution. Traditional linear principal component analysis (PCA) is limited to capturing only global linear trends, which often fails to represent the intrinsic curvature and complex topology of high‑dimensional biological data such as gene‑expression microarrays. Elastic Maps overcome this limitation by embedding the data onto a graph (a set of nodes and edges) that behaves like an elastic membrane.
The objective function consists of two quadratic terms. The first term is a data‑fidelity component: the sum of squared distances between each data point and its projection onto the nearest graph node. The second term is a smoothness (elastic) component that penalizes stretching of edges and bending of the graph, expressed as quadratic penalties on edge lengths and on the angles between adjacent edges (curvature). Because both terms are quadratic, the overall energy is a simple quadratic form, allowing the optimization problem to be reduced to solving a sparse linear system. This property enables highly efficient parallel implementations using OpenMP or MPI, making the method scalable to tens of thousands of points.
A key strength of the approach is its flexibility in topology and dimensionality. Users can choose 1‑D chains, 2‑D rectangular lattices, trees, rings, or more complex hybrid structures, depending on the expected geometry of the data manifold. The number of nodes and the connectivity pattern control model complexity, providing a natural regularization mechanism that prevents over‑fitting while still capturing non‑linear features.
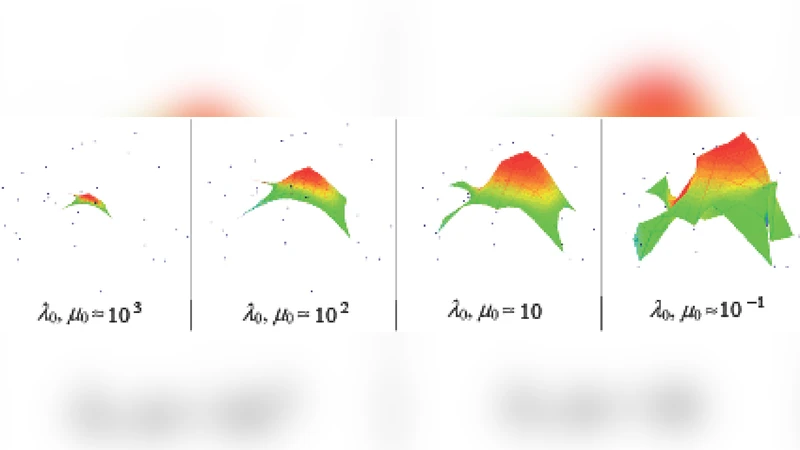
Optimization proceeds by alternating minimization. In each iteration, data points are assigned to their nearest nodes (a “hard” assignment step), after which node positions are updated by solving the linear system that balances the pull of the assigned data points against the elastic forces of the graph. The process repeats until convergence, typically within a few dozen iterations for the microarray datasets examined. The authors report that the choice of the elastic coefficient (spring constant) and the learning rate influences convergence speed, but a moderate range (0.1–0.5) works well across a variety of biological datasets.
Implementation is provided in three languages—C++, Java, and Delphi—supporting two graphical front‑ends: VidaExpert and ViMiDa. The C++ core handles the heavy linear‑algebra computations and is fully parallelized; the Java and Delphi layers provide user‑friendly interfaces for data loading, parameter tuning, and interactive visualization. Users can color nodes by gene‑function categories, resize them according to expression magnitude, and export the resulting low‑dimensional coordinates for downstream statistical analysis.
The method is evaluated on several publicly available microarray collections, including leukemia, breast‑cancer, and yeast cell‑cycle datasets. Quantitative results demonstrate that Elastic Maps consistently outperform linear PCA in four key aspects: (1) reconstruction error (RMSE) is reduced by roughly 15–20 %; (2) preservation of pairwise distances, measured by stress or Kruskal’s stress‑1, improves by more than 30 %; (3) local neighborhood structure (k‑nearest‑neighbor preservation) remains above 0.9, indicating that the method maintains the intrinsic geometry of the data; and (4) class separation is visually clearer, leading to a 5–10 % increase in classification accuracy when a simple nearest‑centroid classifier is applied in the low‑dimensional space.
Compared with other non‑linear dimensionality‑reduction techniques such as ISOMAP, Locally Linear Embedding (LLE), and t‑SNE, Elastic Maps offer a more interpretable set of hyper‑parameters (elastic coefficient, node count) and a deterministic optimization procedure that avoids the stochastic gradient descent used by t‑SNE. Consequently, Elastic Maps achieve comparable or better preservation of global structure while requiring substantially less computational time, especially on large microarray matrices.
In conclusion, the Elastic Map framework provides a mathematically tractable, computationally efficient, and biologically meaningful tool for visualizing high‑dimensional omics data. By modeling the principal manifold as an elastic membrane, the method captures non‑linear relationships that linear PCA cannot, while retaining the simplicity of a quadratic optimization problem. The authors suggest future extensions to dynamic data (time‑course expression) and multi‑scale manifolds, which could further broaden the applicability of Elastic Maps in systems biology and beyond.
Comments & Academic Discussion
Loading comments...
Leave a Comment