Physical approaches to DNA sequencing and detection
With the continued improvement of sequencing technologies, the prospect of genome-based medicine is now at the forefront of scientific research. To realize this potential, however, we need a revolutionary sequencing method for the cost-effective and rapid interrogation of individual genomes. This capability is likely to be provided by a physical approach to probing DNA at the single nucleotide level. This is in sharp contrast to current techniques and instruments which probe, through chemical elongation, electrophoresis, and optical detection, length differences and terminating bases of strands of DNA. In this Colloquium we review several physical approaches to DNA detection that have the potential to deliver fast and low-cost sequencing. Center-fold to these approaches is the concept of nanochannels or nanopores which allow for the spatial confinement of DNA molecules. In addition to their possible impact in medicine and biology, the methods offer ideal test beds to study open scientific issues and challenges in the relatively unexplored area at the interface between solids, liquids, and biomolecules at the nanometer length scale. We emphasize the physics behind these methods and ideas, critically describe their advantages and drawbacks, and discuss future research opportunities in this field.
💡 Research Summary
The paper opens by contrasting the prevailing next‑generation sequencing (NGS) platforms, which rely on chemical polymerisation, electrophoretic separation and optical read‑out, with the emerging need for a truly low‑cost, rapid method capable of interrogating individual genomes. The authors argue that a fundamentally different strategy—probing DNA at the single‑nucleotide level through physical interactions rather than chemistry—offers a path toward that goal. Central to this “physical sequencing” paradigm is the confinement of DNA within nanometer‑scale structures such as nanochannels and nanopores, which translate molecular features into measurable electrical, optical, or mechanical signals.
The review is organized around four principal physical approaches.
-
Nanopore ionic‑current blockade – Biological pores (e.g., α‑hemolysin), solid‑state silicon or graphene pores, and hybrid designs are described. As a single DNA strand threads through a pore, it partially occludes the ionic current flowing through the electrolyte. The magnitude of current reduction depends on the volume, charge, and hydration shell of the nucleotide occupying the constriction. High‑bandwidth current amplifiers can capture these transient events, and statistical models are used to infer base identity. The authors highlight the principal challenges: sub‑picoampere noise, surface charge heterogeneity, and the modest current difference between adjacent bases, which together limit base‑calling accuracy.
-
Nanoconfined optical/electron detection in nanochannels – By fabricating channels with diameters comparable to the DNA double helix (2–5 nm), a strand can be stretched linearly, eliminating the need for fluorescent labels. The confined geometry enables direct interrogation via Raman scattering, plasmonic enhancement, or transmission electron microscopy (TEM) contrast, each of which is sensitive to the electronic structure of individual bases. The review discusses fabrication techniques (focused ion beam, electron‑beam lithography, dielectric breakdown) and the importance of controlling DNA‑channel electrostatic interactions to avoid distortion of the molecule.
-
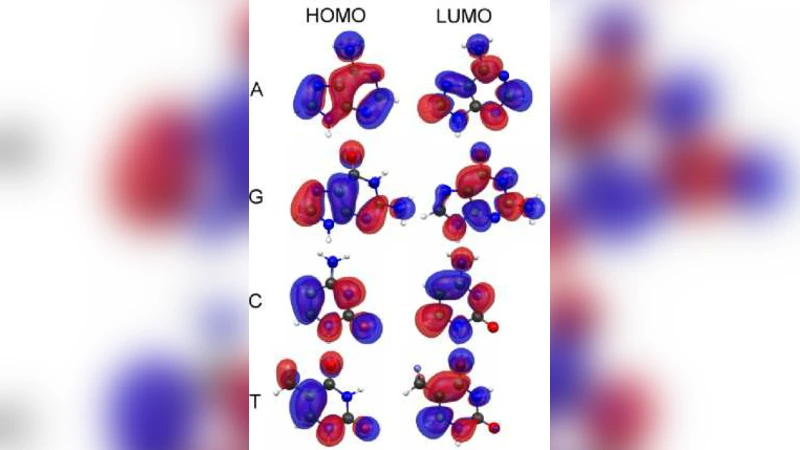
Transverse tunnelling (electron‑transport) measurements – Two nano‑electrodes positioned on opposite sides of a DNA strand can detect the tunnelling current that flows when a bias is applied across the nucleobase. Because tunnelling probability decays exponentially with electrode separation, sub‑nanometre spacing and atomically flat electrode surfaces (often made from gold, carbon nanotubes or graphene) are essential. The current is highly sensitive to the nucleobase’s highest occupied molecular orbital (HOMO) and lowest unoccupied molecular orbital (LUMO) energies, providing a potential electronic “fingerprint.” The authors note that reproducible electrode fabrication, thermal drift, and the need for ultra‑low‑noise amplification remain formidable obstacles.
-
Mechanical force spectroscopy (optical tweezers, AFM) – By pulling a DNA molecule through a nanoscale constriction or directly stretching it with an optical trap or atomic‑force microscope, one can record force‑extension curves. Subtle variations in the force required to unzip or slide each base reflect differences in hydrogen‑bonding and stacking interactions. While this method yields high signal‑to‑noise ratios, its throughput is limited by the serial nature of force measurements and the complexity of deconvolving overlapping mechanical signatures.
Across all approaches, the authors emphasize common advantages: single‑molecule resolution, elimination of costly reagents, and the possibility of real‑time read‑out. Conversely, they identify universal drawbacks: the need for exquisite control of nanoscale geometry, the prevalence of stochastic noise, and the challenge of scaling from proof‑of‑concept devices to high‑throughput sequencers.
The paper concludes with a forward‑looking agenda. Advances in two‑dimensional materials (graphene, MoS₂) promise atomically thin, robust nanopores with tunable surface chemistry. Integration of plasmonic or quantum‑dot sensors could boost optical signal strength, while cryogenic or low‑temperature electronics may suppress thermal noise in tunnelling measurements. Crucially, the authors advocate the use of machine‑learning algorithms to decode complex, noisy signal patterns, turning raw physical data into accurate base calls.
Finally, the authors argue that physical DNA sequencing is not only a potential revolution for genomics and personalized medicine but also a fertile testbed for fundamental studies at the interface of solids, liquids and biomolecules. By confronting the unresolved physics of charge transport, confinement‑induced conformational changes, and nanoscale fluid dynamics, this field will drive interdisciplinary innovation across nanotechnology, biophysics and computational science.
Comments & Academic Discussion
Loading comments...
Leave a Comment