Offdiagonal complexity: A computationally quick network complexity measure. Application to protein networks and cell division
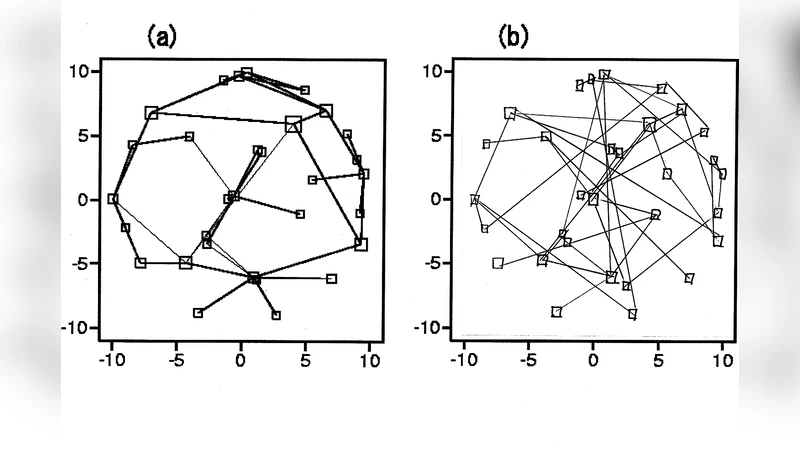
Many complex biological, social, and economical networks show topologies drastically differing from random graphs. But, what is a complex network, i.e.\ how can one quantify the complexity of a graph? Here the Offdiagonal Complexity (OdC), a new, and computationally cheap, measure of complexity is defined, based on the node-node link cross-distribution, whose nondiagonal elements characterize the graph structure beyond link distribution, cluster coefficient and average path length. The OdC apporach is applied to the {\sl Helicobacter pylori} protein interaction network and randomly rewired surrogates thereof. In addition, OdC is used to characterize the spatial complexity of cell aggregates. We investigate the earliest embryo development states of Caenorhabditis elegans. The development states of the premorphogenetic phase are represented by symmetric binary-valued cell connection matrices with dimension growing from 4 to 385. These matrices can be interpreted as adjacency matrix of an undirected graph, or network. The OdC approach allows to describe quantitatively the complexity of the cell aggregate geometry.
💡 Research Summary
The paper introduces Offdiagonal Complexity (OdC), a novel, computationally inexpensive metric for quantifying the structural complexity of graphs. Traditional complexity measures—degree distribution, clustering coefficient, average shortest‑path length—capture only limited aspects of network architecture and often assign similar values to fundamentally different graphs. OdC overcomes this limitation by focusing on the off‑diagonal elements of the node‑node link cross‑distribution, i.e., the frequency with which nodes of different degrees are directly connected.
Formally, for an undirected graph with adjacency matrix A, each node i has degree k_i. By counting how many edges connect a node of degree k to a node of degree ℓ, a two‑dimensional histogram H(k,ℓ) is built. The diagonal entries H(k,k) represent connections between nodes of equal degree, while the off‑diagonal entries (k≠ℓ) encode heterogeneity in degree‑pair connections. OdC is defined as the normalized entropy of these off‑diagonal entries:
OdC = – Σ_{k≠ℓ}
Comments & Academic Discussion
Loading comments...
Leave a Comment