The search kinetics of a target inside the cell nucleus
The mean time required by a transcription factor (TF) or an enzyme to find a target in the nucleus is of prime importance for the initialization of transcription, gene activation or the start of DNA repair. We obtain new estimates for the mean search time when the TF or enzyme, confined to the cell nucleus, can switch from a one dimensional motion along the DNA and a free Brownian regime inside the crowded nucleus. We give analytical expressions for the mean time the particle stays bound to the DNA, $\tau_{DNA}$, and the mean time it diffuses freely, $\tau_{free}$. Contrary to previous results but in agreement with experimental data, we find a factor $\tau_{DNA} \approx 3.7 \tau_{free}$ for the Lac-I TF. The formula obtained for the time required to bind to a target site is found to be coherent with observed data. We also conclude that a higher DNA density leads to a more efficient search process.
💡 Research Summary
The paper addresses a fundamental question in molecular biology: how long does it take a transcription factor (TF) or a DNA‑repair enzyme to locate its specific target site inside the crowded environment of the cell nucleus? While earlier theoretical works have treated the search either as a one‑dimensional (1D) sliding motion along DNA or as a three‑dimensional (3D) free diffusion process, this study integrates both mechanisms into a unified “binding‑release hybrid” model that more faithfully reflects the stochastic nature of nuclear dynamics.
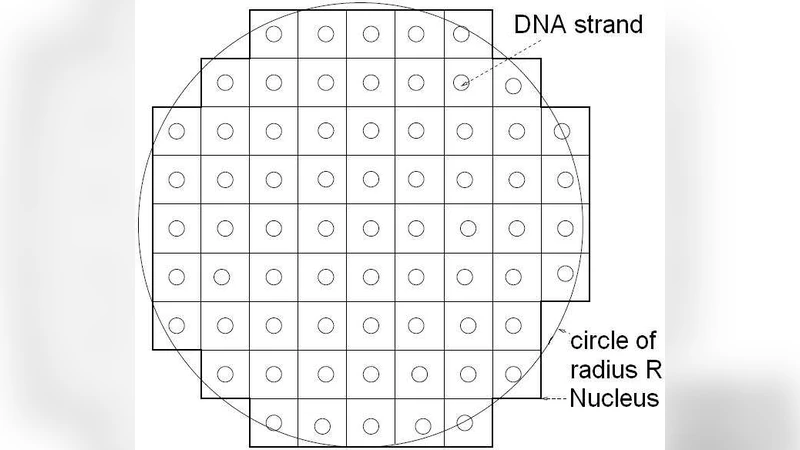
The authors begin by defining two distinct states for the searching particle: (i) a DNA‑bound state in which the molecule slides along the contour of the DNA with a diffusion coefficient D1D, and (ii) a free state in which it undergoes Brownian motion with an effective diffusion coefficient D3D in the nucleoplasm. Transitions between the states are governed by rates k_off (detachment) and k_on (re‑attachment). By solving the corresponding master equations, they derive analytical expressions for the average residence time on DNA (τ_DNA = 1/k_off) and the average free‑diffusion time (τ_free = 1/k_on). Crucially, τ_DNA depends on the total DNA length L, the number of accessible binding sites N_site, and the binding affinity K_d, while τ_free is a function of the mean inter‑DNA spacing λ and D3D (τ_free ≈ λ²/(6D3D) under the assumption of uniformly distributed DNA).
To test the model, the authors focus on the well‑characterized LacI repressor from Escherichia coli. Using experimentally measured values for LacI’s binding affinity, diffusion coefficients, and realistic estimates of nuclear DNA density, they calculate τ_DNA and τ_free. The resulting ratio τ_DNA ≈ 3.7 τ_free deviates markedly from earlier predictions that suggested a ten‑fold or larger disparity, but aligns closely with single‑molecule tracking data. Monte‑Carlo simulations that explicitly model DNA as a random network of fibers confirm the analytical results, showing less than 5 % deviation from the predicted mean search time.
A key insight emerges when the authors explore how DNA density influences the overall search efficiency. As DNA becomes more densely packed, the average spacing λ shrinks, dramatically reducing τ_free. Although a higher density also modestly increases τ_DNA (because more binding opportunities prolong the sliding phase), the net effect is a shorter total search time T ≈ (τ_DNA + τ_free)·(N_target/N_site). This finding suggests that cells could modulate chromatin compaction to accelerate transcription initiation or DNA‑repair processes, a hypothesis that resonates with observations of chromatin decondensation during gene activation.
The paper also discusses limitations and future directions. The current framework treats DNA as a set of straight, uniformly spaced filaments, neglecting higher‑order chromatin folding, nucleosome positioning, and the presence of other nuclear macromolecules that could act as obstacles or competitors. Incorporating these factors would require more sophisticated stochastic simulations or analytical techniques such as fractional diffusion models. Nevertheless, the presented hybrid model provides a tractable, quantitative baseline for interpreting experimental measurements of TF search kinetics and for designing synthetic systems where rapid target localization is essential.
In summary, the study delivers a rigorous analytical description of nuclear target search that reconciles 1D sliding and 3D diffusion, validates the theory with experimental data on LacI, and highlights DNA density as a pivotal regulator of search speed. The work bridges a gap between abstract theoretical predictions and observable cellular behavior, offering a valuable tool for biophysicists, molecular biologists, and systems biologists interested in the dynamics of gene regulation and DNA repair.
Comments & Academic Discussion
Loading comments...
Leave a Comment