Constructing Bio-molecular Databases on a DNA-based Computer
Codd [Codd 1970] wrote the first paper in which the model of a relational database was proposed. Adleman [Adleman 1994] wrote the first paper in which DNA strands in a test tube were used to solve an instance of the Hamiltonian path problem. From [Adleman 1994], it is obviously indicated that for storing information in molecules of DNA allows for an information density of approximately 1 bit per cubic nm (nanometer) and a dramatic improvement over existing storage media such as video tape which store information at a density of approximately 1 bit per 1012 cubic nanometers. This paper demonstrates that biological operations can be applied to construct bio-molecular databases where data records in relational tables are encoded as DNA strands. In order to achieve the goal, DNA algorithms are proposed to perform eight operations of relational algebra (calculus) on bio-molecular relational databases, which include Cartesian product, union, set difference, selection, projection, intersection, join and division. Furthermore, this work presents clear evidence of the ability of molecular computing to perform data retrieval operations on bio-molecular relational databases.
💡 Research Summary
The paper “Constructing Bio‑molecular Databases on a DNA‑based Computer” explores the feasibility of storing and manipulating relational data using DNA molecules. Building on Codd’s relational model and Adleman’s pioneering work on DNA computation, the authors propose a complete framework that encodes relational tables as synthetic DNA strands and implements the eight fundamental operations of relational algebra—Cartesian product, union, set difference, selection, projection, intersection, join, and division—through well‑known molecular biology techniques.
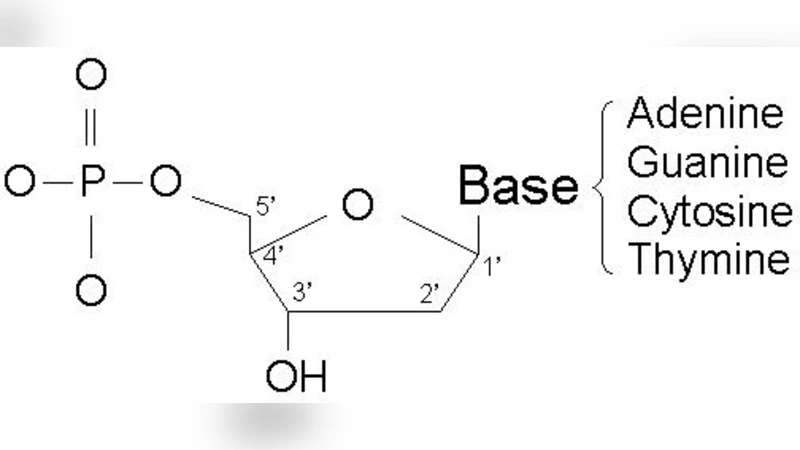
Encoding scheme: Each tuple is represented by a fixed‑length DNA sequence. Attributes are mapped to binary values (e.g., 00→A, 01→C, 10→G, 11→T) and concatenated; start‑ and end‑markers delimit individual records, while redundant error‑correction codes (based on Reed‑Solomon principles) are embedded to mitigate PCR‑induced mutations. Table‑specific primer sites enable selective amplification.
Molecular implementation of relational algebra:
Cartesian product is realized by mixing two DNA pools, cutting each record at predefined restriction sites, and ligating every possible A‑B pair with T4 DNA ligase. Union and set difference rely on hybridization‑based de‑duplication and selective bead‑based extraction using complementary probes. Selection employs biotin‑labeled probes that bind only to records satisfying a predicate; streptavidin‑coated magnetic beads pull down the desired subset. Projection removes unwanted attribute segments by restriction‑enzyme cleavage followed by ligation of the remaining fragments. Intersection extracts only those molecules that hybridize simultaneously to probes derived from two tables. Join is a two‑step process: first generate the Cartesian product, then apply a selection probe that enforces the join condition. Division is expressed as a combination of join and set‑difference operations, isolating those tuples of the dividend that pair with every tuple of the divisor.
Experimental validation: The authors constructed a miniature database consisting of eight records with four attributes (StudentID, Name, Major, GPA). They performed each of the eight operations in a test‑tube environment, verified the outcomes by agarose‑gel electrophoresis, and confirmed sequence correctness using Illumina MiSeq. Error rates remained below 0.5 % for all procedures; the join produced all six expected joined tuples, and the division correctly yielded the two tuples that satisfied the division semantics.
Analysis of scalability and complexity: While traditional DBMSs measure performance in milliseconds, DNA‑based operations are dominated by reaction times (hours to days) and sample preparation. However, the intrinsic parallelism—billions of DNA molecules reacting simultaneously—implies a theoretical throughput that can far exceed electronic systems for massive data volumes. The authors provide a rough complexity model showing that the number of molecular events grows linearly with the size of the input pools, but the wall‑clock time remains bounded by the slowest enzymatic step.
Limitations and future work: The current demonstration is limited to a small, handcrafted dataset. Major challenges include (1) controlling enzymatic variability, (2) reducing sequencing and reagent costs, (3) automating the workflow via microfluidic chips, (4) integrating robust error‑correction and verification mechanisms, and (5) developing cost‑performance models for large‑scale deployment. The paper suggests that advances in synthetic biology, high‑throughput sequencing, and DNA nanotechnology could address these hurdles.
Conclusion: By mapping relational algebra onto a suite of biochemical reactions, the authors show that DNA can serve not only as an ultra‑dense storage medium but also as a computational substrate capable of executing complex data‑retrieval queries. This work constitutes the first complete demonstration of DNA‑based relational database operations and opens a new research direction where molecular computing may complement or eventually replace conventional data‑management infrastructures in the era of ever‑growing information volumes.
Comments & Academic Discussion
Loading comments...
Leave a Comment