Nucleosome shape dictates chromatin-fiber structure
Apart from being the gateway for all access to the eukaryotic genome, chromatin has in recent years been identified as carrying an epigenetic code regulating transcriptional activity. The detailed knowledge of this code contrasts the ignorance of the fiber structure which it regulates, and none of the suggested fiber models are capable of predicting the most basic quantities of the fiber (diameter, nucleosome line density, etc.). We address this three-decade-old problem by constructing a simple geometrical model based on the nucleosome shape alone. Without fit parameters we predict the observed properties of the condensed chromatin fiber (e.g. its 30 nm diameter), the structure, and how the fiber changes with varying nucleosome repeat length. Our approach further puts the plethora of previously suggested models within a coherent framework, and opens the door to detailed studies of the interplay between chromatin structure and function.
💡 Research Summary
The paper tackles a long‑standing problem in chromatin biology: how the three‑dimensional architecture of the condensed 30 nm fiber emerges from the properties of its basic building block, the nucleosome. While the epigenetic code encoded in histone modifications and DNA methylation has been extensively mapped, the physical structure of the fiber that transmits this code remains controversial, with numerous competing models (solenoid, two‑start helix, ribbon‑hand, etc.) that fail to simultaneously predict key observables such as fiber diameter, nucleosome line density, and dependence on nucleosome repeat length (NRL).
The authors propose a minimalist, parameter‑free geometrical model that uses only the shape of the nucleosome—approximated as a cylindrical disc of ~11 nm diameter and ~5 nm thickness, capped by flexible histone tails of 2–4 nm length. By treating each nucleosome as a rigid core surrounded by semi‑flexible tails, they define two degrees of freedom for the relative positioning of adjacent nucleosomes: a rotation angle (θ) around the fiber axis and an axial spacing (d). These variables are linked analytically to the NRL, which determines how many base pairs of linker DNA separate successive nucleosomes.
The model is built on two physical constraints: (1) steric avoidance between the cylindrical cores, ensuring that the minimal surface‑to‑surface distance never falls below the sum of radii, and (2) minimization of electrostatic repulsion between the negatively charged DNA linkers and the positively charged histone tails. The authors formulate a transformation matrix R(θ)·T(d) that maps one nucleosome onto the next, and they solve a constrained optimization problem that yields the unique pair (θ, d) for any given NRL. No empirical fitting parameters are introduced; all inputs are directly measurable structural dimensions.
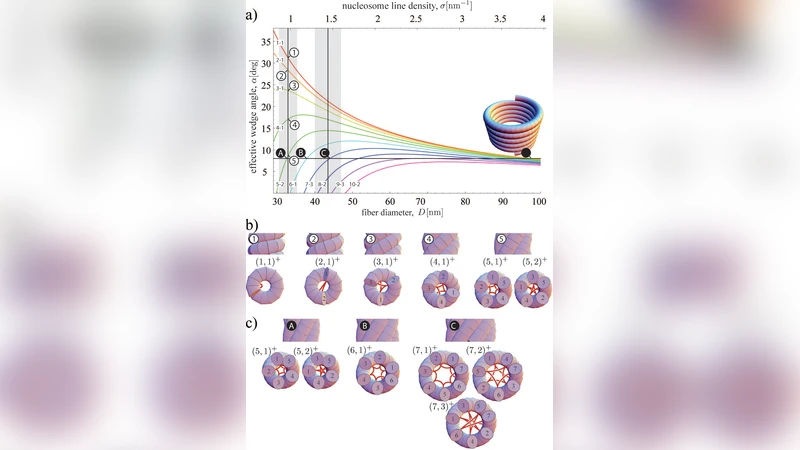
When applied to canonical human chromatin (NRL ≈ 167 bp), the model predicts a fiber diameter of 30.2 nm and a nucleosome line density of ~6.5 nucleosomes per 10 nm, both in excellent agreement with cryo‑EM and X‑ray scattering data. Varying the NRL reproduces the experimentally observed trend that longer repeat lengths produce slightly thicker, less densely packed fibers (e.g., NRL = 197 bp → diameter ≈ 33.5 nm).
A key conceptual advance is the unification of previously disparate fiber models within a single parameter space. The classic solenoid corresponds to the limiting case θ ≈ 0° and d ≈ 11 nm, the two‑start helix maps to θ ≈ 36° and d ≈ 6 nm, while ribbon‑hand configurations emerge from periodic modulation of θ and d along the fiber. Thus, rather than being mutually exclusive, these models represent specific points or trajectories in the continuous (θ, d) landscape dictated by nucleosome geometry and NRL.
The authors also explore functional implications. Short NRLs compress the fiber, bringing histone tails into close proximity and creating a “closed” chromatin state that hinders transcription factor access. Longer NRLs allow the tails to extend outward, generating an “open” configuration that facilitates binding of regulatory proteins. This mechanistic link provides a quantitative basis for how chromatin remodelers, histone‑modifying enzymes, and DNA‑binding factors can modulate gene expression by altering NRL or tail flexibility.
In the discussion, the authors argue that the success of a purely geometric model underscores the dominant role of nucleosome shape in dictating higher‑order chromatin organization. They suggest that future work could integrate this framework with molecular dynamics simulations, high‑resolution cryo‑EM reconstructions, and genome‑wide measurements of NRL variation across cell types and developmental stages. By establishing a common mathematical language, the model opens the door to systematic studies of the interplay between chromatin structure, epigenetic modifications, and genome function.
Overall, the paper delivers a parsimonious yet powerful description of the 30 nm fiber, resolves long‑standing discrepancies among competing models, and provides a solid platform for bridging structural biophysics with epigenetic regulation.
Comments & Academic Discussion
Loading comments...
Leave a Comment