An Empirical Study of Immune System Based On Bipartite Network
Immune system is the most important defense system to resist human pathogens. In this paper we present an immune model with bipartite graphs theory. We collect data through COPE database and construct an immune cell- mediators network. The act degree distribution of this network is proved to be power-law, with index of 1.8. From our analysis, we found that some mediators with high degree are very important mediators in the process of regulating immune activity, such as TNF-alpha, IL-8, TNF-alpha receptors, CCL5, IL-6, IL-2 receptors, TNF-beta receptors, TNF-beta, IL-4 receptors, IL-1 beta, CD54 and so on. These mediators are important in immune system to regulate their activity. We also found that the assortative of the immune system is -0.27. It reveals that our immune system is non-social network. Finally we found similarity of the network is 0.13. Each two cells are similar to small extent. It reveals that many cells have its unique features. The results show that this model could describe the immune system comprehensive.
💡 Research Summary
The paper proposes a quantitative representation of the human immune system using bipartite network theory. By extracting information from the COPE (Comprehensive Protein‑Expression) database, the authors construct a two‑mode graph in which one set of nodes corresponds to immune cells (e.g., T‑cells, B‑cells, macrophages) and the other set to soluble mediators and receptors (cytokines, chemokines, surface receptors). An edge is drawn whenever a cell secretes a mediator or expresses a receptor that can bind that mediator. This formulation captures the many‑to‑many relationships that underlie immune signaling and allows the application of standard network‑analytic tools.
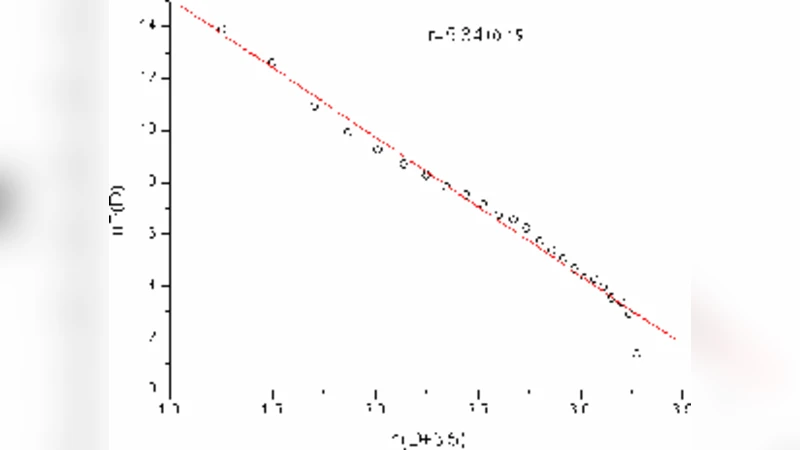
The first major result concerns the degree distribution of the “act” nodes (mediators). The authors find that the distribution follows a power‑law with an exponent of approximately 1.8, indicating a scale‑free structure in which a small number of mediators act as hubs linking many cell types. The identified high‑degree mediators include TNF‑α, IL‑8, CCL5, IL‑6, IL‑2 receptors, TNF‑β receptors, TNF‑β, IL‑4 receptors, IL‑1β, and CD54. These molecules are well known to be central in inflammation, cell activation, and immune regulation, thereby providing biological validation for the network‑based approach.
Next, the authors compute the assortativity coefficient of the bipartite graph, obtaining a value of –0.27. A negative assortativity indicates a disassortative (non‑social) network: high‑degree nodes preferentially connect to low‑degree nodes. In the immune context this reflects the fact that a few broadly acting cytokines interact with many specialized cell types, while most mediators have a narrow target spectrum. This pattern contrasts with many social networks, where high‑degree individuals tend to associate with each other, and underscores the functional organization of immune signaling.
The paper also evaluates pairwise similarity between cells using a Jaccard‑type index based on shared mediators. The average similarity is reported as 0.13, a relatively low value that suggests most immune cells possess distinct mediator repertoires. This finding aligns with the known specialization of immune subsets (e.g., cytotoxic T‑cells versus regulatory T‑cells) and supports the view that the immune system balances diversity with a set of common regulatory hubs.
While the study offers an elegant structural portrait, several methodological limitations merit discussion. First, COPE provides static expression snapshots; immune responses are highly dynamic, with cytokine levels fluctuating over minutes to days. Incorporating temporal data or longitudinal sampling would enable the construction of dynamic bipartite networks that capture activation waves and feedback loops. Second, the projection of the bipartite graph onto a single mode (cell‑cell or mediator‑mediator) inevitably discards information about the bipartite nature of the interactions. The authors do not explore how such projections affect community detection or functional module identification. Third, the paper stops short of linking network metrics to functional outcomes. For instance, does the hub status of TNF‑α correlate with disease severity in sepsis, or can the low similarity index predict susceptibility to autoimmune disorders? Experimental validation—such as perturbation experiments that remove a hub cytokine and observe network rewiring—would strengthen the causal interpretation.
Future work could address these gaps by (a) integrating time‑resolved transcriptomic or proteomic datasets, (b) employing multilayer network frameworks that simultaneously model cell‑cell contact, soluble mediator exchange, and intracellular signaling pathways, and (c) testing predictive models derived from network topology against clinical phenotypes. Moreover, extending the analysis to disease‑specific cohorts (e.g., patients with rheumatoid arthritis, COVID‑19, or cancer immunotherapy) would reveal how the bipartite structure remodels under pathological conditions and might uncover novel therapeutic targets.
In summary, the authors successfully demonstrate that a bipartite network built from cell‑mediator interactions exhibits hallmark features of complex biological systems: a scale‑free degree distribution, disassortative mixing, and low pairwise similarity. These structural characteristics provide a concise, quantitative language for describing immune system organization and lay the groundwork for more sophisticated, dynamic, and disease‑focused network models.
Comments & Academic Discussion
Loading comments...
Leave a Comment