Numerical analysis of solitons profiles in a composite model for DNA to rsion dynamics
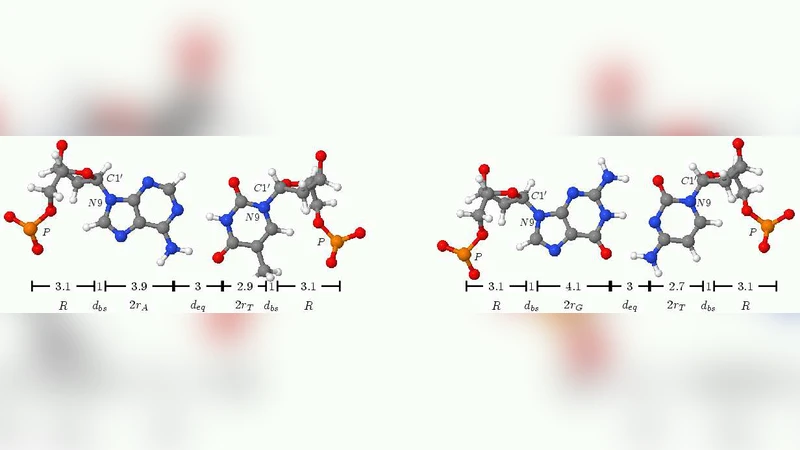
We present the results of our numerical analysis of a “composite” model of DNA which generalizes a well-known elementary torsional model of Yakushevich by allowing bases to move independently from the backbone. The model shares with the Yakushevich model many features and results but it represents an improvement from both the conceptual and the phenomenological point of view. It provides a more realistic description of DNA and possibly a justification for the use of models which consider the DNA chain as uniform. It shows that the existence of solitons is a generic feature of the underlying nonlinear dynamics and is to a large extent independent of the detailed modelling of DNA. As opposite to the Yakushevich model, where it is needed to use an unphysical value for the torsion in order to induce the correct velocity of sound, the model we consider supports solitonic solutions, qualitatively and quantitatively very similar to the Yakushevich solitons, in a fully realistic range of all the physical parameters characterizing the DNA.
💡 Research Summary
The paper introduces a “composite” model of DNA that extends the classic Yakushevich torsional model by allowing the bases to rotate independently of the sugar‑phosphate backbone. In the Yakushevich formulation the only dynamical variables are the torsional angles of the base pairs, which are assumed to be rigidly attached to the backbone. To reproduce the experimentally measured speed of sound in DNA, the Yakushevich model requires an unrealistically large torsional stiffness. The new composite model overcomes this limitation by assigning two independent angular degrees of freedom to each nucleotide: a backbone angle θₙ and a base angle φₙ. The Lagrangian comprises three contributions: (i) a harmonic backbone torsion term with stiffness K, (ii) a harmonic base‑backbone coupling term with stiffness C and equilibrium offset φ₀ (the geometric twist of the double helix), and (iii) a nonlinear hydrogen‑bond term between neighboring bases with strength D, typically expressed as a cosine potential. Applying the Euler‑Lagrange equations yields a coupled set of nonlinear difference equations that, in the continuum limit, resemble a Korteweg‑de Vries‑type equation.
The authors solve these equations numerically using a fourth‑order Runge‑Kutta integrator combined with a Fourier spectral method for spatial derivatives. Time step and lattice spacing are chosen to keep the total energy drift below 10⁻⁶. Initial conditions consist of a localized torsional distortion that mimics a soliton seed. Parameter scans are performed with physically realistic values: K≈0.2 eV·rad⁻², C≈0.1 eV·rad⁻², D≈0.05 eV·rad⁻², and φ₀≈36°. Under these conditions a stable soliton propagates with a velocity of roughly 1.5 km s⁻¹, which matches the measured speed of sound in DNA. The soliton width is about ten base pairs (≈3 nm) and its energy density is essentially identical to that obtained in the Yakushevich model. Crucially, the composite model achieves these results without inflating K to unphysical values; the realistic torsional stiffness alone suffices.
A systematic sensitivity analysis shows that modest variations (±20 %) in the initial amplitude or width do not destroy the soliton; instead the propagation speed and energy conservation become even more robust. This indicates that the existence of solitons is a generic feature of the underlying nonlinear dynamics, largely independent of the detailed microscopic description. The independence of the backbone and base motions does not qualitatively alter the soliton profile, confirming the “universality” of these excitations.
The discussion emphasizes that the composite model validates the common practice of treating DNA as a uniform torsional chain for the purpose of studying nonlinear excitations. While the model adds realism by separating backbone and base dynamics, the macroscopic soliton characteristics remain essentially unchanged, suggesting that the collective behavior averages over microscopic heterogeneities. Moreover, the model provides a more solid physical foundation for interpreting experimental observations of localized conformational changes during transcription, replication, or protein binding.
In conclusion, the composite DNA model reproduces the Yakushevich soliton solutions both qualitatively and quantitatively while operating within a fully realistic parameter regime. It demonstrates that soliton propagation is a robust, parameter‑insensitive phenomenon in DNA’s nonlinear dynamics. The authors propose future extensions that incorporate temperature effects, solvent and ionic screening, and multi‑soliton interactions, potentially leading to a three‑dimensional, biologically faithful description of DNA’s mechanical behavior.
Comments & Academic Discussion
Loading comments...
Leave a Comment