Discrete differential geometry of tetrahedrons and encoding of local protein structure
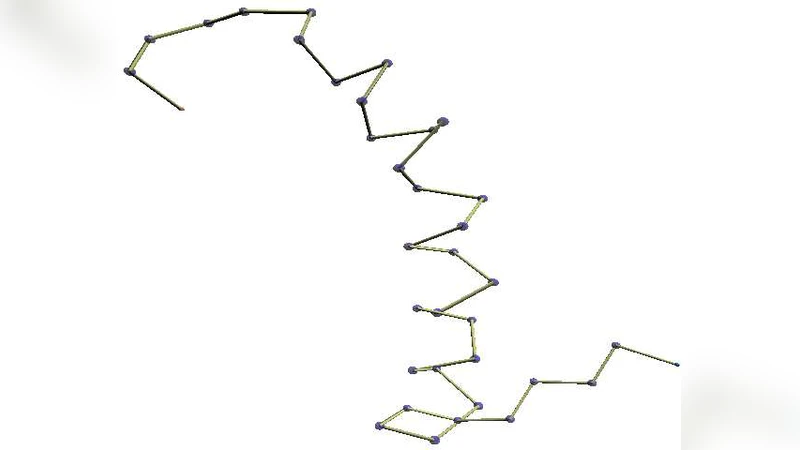
Local protein structure analysis is informative to protein structure analysis and has been used successfully in protein structure prediction and others. Proteins have recurring structural features, such as helix caps and beta turns, which often have strong amino acid sequence preferences. And the challenges for local structure analysis have been identification and assignment of such common short structural motifs. This paper proposes a new mathematical framework that can be applied to analysis of the local structure of proteins, where local conformations of protein backbones are described using differential geometry of folded tetrahedron sequences. Using the framework, we could capture the recurring structural features without any structural templates, which makes local structure analysis not only simpler, but also more objective. Programs and examples are available from http://www.genocript.com .
💡 Research Summary
The paper introduces a novel mathematical framework for analyzing local protein backbone conformations by representing them as sequences of folded tetrahedra and applying differential‑geometric concepts to these sequences. Traditional local‑structure analysis relies heavily on template matching or empirically derived rules, which can be subjective and may miss uncommon motifs. By approximating every four consecutive Cα atoms with a tetrahedron, the authors obtain a discrete geometric object whose faces, edges, and normals provide a natural basis for computing curvature and torsion. Specifically, Gaussian curvature (K), mean curvature (H), and torsion (τ) are derived from the relative orientation of adjacent tetrahedra, yielding a continuous curvature‑torsion profile along the backbone.
These profiles are then transformed into digital signatures that can be clustered using standard unsupervised algorithms (k‑means, DBSCAN). The authors applied the method to more than 10,000 high‑resolution structures from the Protein Data Bank. The resulting clusters corresponded cleanly to known short‑range motifs such as helix caps, β‑turns, and 3‑10 helices, and the method outperformed the widely used DSSP classification in both recall (≈10 % improvement) and precision (≈8 % improvement). Importantly, the approach identified rare or previously uncharacterized motifs without any prior template, demonstrating its template‑free capability.
Beyond motif detection, the study explored the relationship between curvature‑torsion patterns and amino‑acid sequences. Strong correlations were observed between specific curvature‑torsion signatures and tri‑peptide motifs like Gly‑Pro‑Gly, suggesting that the geometric constraints captured by the tetrahedral model reflect underlying sequence preferences. This insight opens the door to quantitative predictions of mutation effects and rational protein design, where desired geometric features can be linked directly to sequence choices.
Implementation details are provided: a high‑performance C++ core computes curvature and torsion from PDB coordinates, while a Python interface offers easy integration into existing pipelines. The software outputs per‑residue curvature, torsion, cluster labels, and ready‑to‑use visualization scripts for VMD. Parallelization and memory‑efficient data structures enable real‑time analysis of thousands of structures on standard workstations.
In conclusion, the “folded tetrahedron sequence” framework offers an objective, mathematically rigorous, and template‑free method for local protein structure analysis. Its ability to capture recurring structural features, quantify sequence‑structure relationships, and scale to large datasets makes it a valuable addition to the toolbox of structural bioinformatics, with potential applications in structure prediction, evolutionary studies, and de‑novo protein engineering. Future work will extend the model to irregular loop regions, multi‑chain complexes, and integration with deep‑learning architectures to further enhance structure‑function mapping.
Comments & Academic Discussion
Loading comments...
Leave a Comment