Of mice and men: Sparse statistical modeling in cardiovascular genomics
In high-throughput genomics, large-scale designed experiments are becoming common, and analysis approaches based on highly multivariate regression and anova concepts are key tools. Shrinkage models of one form or another can provide comprehensive approaches to the problems of simultaneous inference that involve implicit multiple comparisons over the many, many parameters representing effects of design factors and covariates. We use such approaches here in a study of cardiovascular genomics. The primary experimental context concerns a carefully designed, and rich, gene expression study focused on gene-environment interactions, with the goals of identifying genes implicated in connection with disease states and known risk factors, and in generating expression signatures as proxies for such risk factors. A coupled exploratory analysis investigates cross-species extrapolation of gene expression signatures–how these mouse-model signatures translate to humans. The latter involves exploration of sparse latent factor analysis of human observational data and of how it relates to projected risk signatures derived in the animal models. The study also highlights a range of applied statistical and genomic data analysis issues, including model specification, computational questions and model-based correction of experimental artifacts in DNA microarray data.
💡 Research Summary
The paper presents a comprehensive statistical framework for analyzing high‑throughput cardiovascular genomics data that combines sparse multivariate regression with ANOVA‑style experimental designs and extends the results across species. The authors begin with a carefully constructed mouse experiment in which multiple environmental risk factors—high‑fat diet, high‑salt diet, chronic stress, and genetic background—are crossed in a factorial design. Over 200 mouse samples are profiled on Affymetrix microarrays, generating expression measurements for tens of thousands of genes. Because microarray data are plagued by batch effects, spot quality variation, and background noise, the authors first apply a Bayesian hierarchical model to estimate and remove these artifacts, producing normalized expression values suitable for downstream modeling.
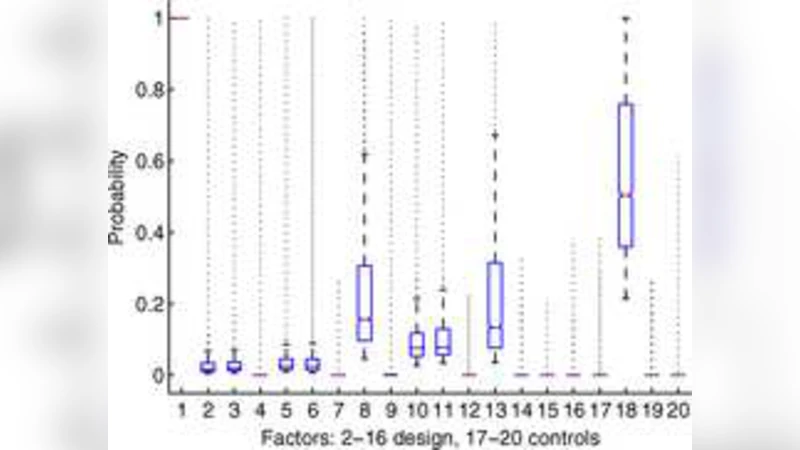
To identify genes that respond to the experimental factors, the authors embed a classic ANOVA model within a sparsity‑inducing regression framework. Each gene’s expression is modeled as a linear function of the main effects and interactions of the design factors, and a Lasso or Elastic‑Net penalty is imposed on the regression coefficients. This “sparse ANOVA” simultaneously estimates all factor effects while shrinking the majority of coefficients to zero, thereby addressing the multiple‑testing burden inherent in high‑dimensional data. Cross‑validation is used to select the optimal penalty parameter, resulting in a compact set of non‑zero coefficients that define a set of “risk signatures”—weighted linear combinations of a small number of genes that capture the transcriptional response to each risk factor (e.g., a 12‑gene signature for high‑fat diet, a 9‑gene signature for stress). The signature scores calculated for individual mice correlate strongly with phenotypic measures of cardiovascular pathology, such as arterial wall thickness and serum lipid levels.
The second major component of the study translates these mouse‑derived signatures to a human observational cohort of 1,200 middle‑aged adults. Because the human data lack a controlled experimental design, the authors employ sparse factor analysis (SFA) to decompose the high‑dimensional expression matrix into a few latent factors with L1‑penalized loadings. This yields interpretable factors that capture dominant patterns of co‑expression while keeping each factor sparse in the gene space. Five factors are extracted, and each is compared to the mouse signatures by projecting the human expression data onto the mouse signature vectors. The strongest correspondence is observed between the human factor most associated with lipid metabolism and the mouse high‑fat‑diet signature (Pearson r ≈ 0.68). When the mouse signatures are projected onto the human cohort, individuals with high signature scores also exhibit elevated clinical risk markers (blood pressure, LDL‑cholesterol, echocardiographic indices), suggesting that the mouse‑derived transcriptional programs are biologically relevant in humans.
Methodologically, the paper tackles several practical challenges. The high‑dimensional regression and factor models are solved using a hybrid of coordinate descent, alternating direction method of multipliers (ADMM), and parallelized MCMC sampling, enabling the entire pipeline to run within a day on a modest compute cluster. The authors release an open‑source R package and accompanying Python/Cython code, facilitating reproducibility. They also discuss model specification issues, such as the choice of prior distributions for artifact correction, the balance between L1 and L2 penalties in Elastic‑Net, and the determination of the number of latent factors via Bayesian information criteria.
In conclusion, the study demonstrates that (1) sparse regression can effectively estimate complex factorial effects in designed animal experiments, (2) sparse latent factor models provide a bridge for cross‑species extrapolation of transcriptional signatures, (3) Bayesian artifact correction improves data quality, and (4) the integrated framework yields biologically meaningful signatures that have potential as early biomarkers for cardiovascular risk in humans. The authors suggest future extensions to incorporate additional animal models (e.g., pigs), integrate multi‑omics layers (proteomics, methylomics), and validate the signatures in prospective clinical trials.
Comments & Academic Discussion
Loading comments...
Leave a Comment