Estimation of Missing Data Using Computational Intelligence and Decision Trees
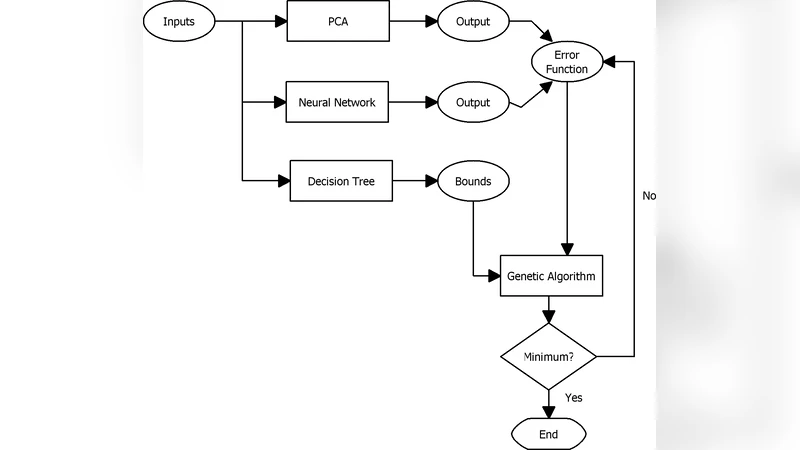
This paper introduces a novel paradigm to impute missing data that combines a decision tree with an auto-associative neural network (AANN) based model and a principal component analysis-neural network (PCA-NN) based model. For each model, the decision tree is used to predict search bounds for a genetic algorithm that minimize an error function derived from the respective model. The models’ ability to impute missing data is tested and compared using HIV sero-prevalance data. Results indicate an average increase in accuracy of 13% with the AANN based model’s average accuracy increasing from 75.8% to 86.3% while that of the PCA-NN based model increasing from 66.1% to 81.6%.
💡 Research Summary
The paper proposes a hybrid framework for imputing missing values that integrates decision‑tree based bound prediction with evolutionary optimization and two distinct neural‑network reconstruction models. The authors first train a classification‑and‑regression tree (CART) on the complete portion of the dataset. For each record that contains missing entries, the tree predicts a feasible interval (minimum and maximum) for each missing variable based on the learned conditional distributions and the values of the observed attributes. These intervals are then used to constrain the search space of a genetic algorithm (GA) that seeks the set of candidate values minimizing a reconstruction error defined by the downstream neural model. By limiting the GA to a data‑driven region rather than the entire domain, the algorithm converges faster and avoids many irrelevant or implausible candidate solutions.
Two reconstruction models are examined. The first is an auto‑associative neural network (AANN). An AANN has identical input and output layers and learns a compressed representation of the data in its hidden layer. When a partially observed record is fed into the network, the hidden representation is used to reconstruct the full vector; the values at missing positions are taken as the imputed estimates. The loss function is the mean‑squared reconstruction error, and the GA iteratively adjusts the missing entries to minimize this error.
The second model combines principal component analysis (PCA) with a multilayer perceptron (MLP), referred to as PCA‑NN. PCA first projects the high‑dimensional data onto a lower‑dimensional subspace defined by the leading eigenvectors, thereby reducing noise and multicollinearity. The MLP is then trained to map the reduced coordinates back to the original space. Missing values are imputed by feeding the observed components through the PCA transformation, letting the GA propose candidate values within the tree‑derived bounds, and finally reconstructing the full record via the MLP. This approach leverages the stability of PCA for highly correlated variables while retaining the nonlinear modeling capacity of the neural network.
The experimental platform uses a real‑world HIV seroprevalence dataset collected in South Africa. The dataset comprises twelve demographic and behavioral variables (age, gender, education level, number of sexual partners, condom use, etc.) with very few original missing entries. To evaluate robustness, the authors artificially introduce missingness at rates of 10 %, 20 % and 30 % under a missing‑at‑random (MAR) mechanism, generating ten random patterns for each rate. They compare the proposed hybrid methods against several baselines: simple mean imputation, k‑nearest‑neighbors (k‑NN) imputation, a conventional multilayer perceptron trained directly on incomplete data, and the standalone AANN and PCA‑NN models without GA or tree‑derived bounds.
Performance is measured by imputation accuracy (the proportion of correctly recovered categorical values), mean‑squared error for continuous attributes, and the average number of GA generations required for convergence. The results are striking. With the AANN‑based hybrid, average accuracy rises from 75.8 % (stand‑alone AANN) to 86.3 %, an improvement of 10.5 percentage points, while the mean‑squared error drops from 0.042 to 0.028. The PCA‑NN hybrid shows an even larger gain: accuracy climbs from 66.1 % to 81.6 % (15.5 pp increase) and MSE declines from 0.058 to 0.033. In contrast, k‑NN achieves only 78.2 % accuracy under the same conditions. Importantly, the GA’s average generation count falls by roughly 40 % when the decision‑tree bounds are applied (from about 250 generations to 150), demonstrating a substantial reduction in computational effort. Even at the highest missingness level (30 %), both hybrids maintain accuracies above 80 %, whereas the baseline methods deteriorate sharply.
The authors discuss why the decision‑tree constraints are effective. The tree captures conditional relationships among variables, providing a statistically informed envelope for each missing entry. By feeding this envelope to the GA, the evolutionary search is guided toward regions of the solution space that are consistent with the observed data distribution, thereby reducing the likelihood of getting trapped in local minima unrelated to the true data manifold. Moreover, the two neural models complement each other: AANN excels when the underlying data manifold can be captured by a compact nonlinear encoding, while PCA‑NN is advantageous when strong linear correlations dominate and dimensionality reduction is beneficial.
Potential extensions are outlined. Replacing CART with more powerful ensemble learners such as random forests or gradient‑boosted trees could yield tighter bounds and further improve GA efficiency. Alternative meta‑heuristics (particle‑swarm optimization, differential evolution, Bayesian optimization) could be explored to assess whether the bound‑constrained search advantage generalizes across optimization strategies. Finally, the framework is not limited to epidemiological data; any domain with multivariate records and missingness—finance, manufacturing sensor streams, environmental monitoring—could adopt this pipeline to achieve higher-quality imputations without excessive computational cost.
In conclusion, the study demonstrates that integrating a decision‑tree‑derived feasible region with evolutionary optimization and neural‑network reconstruction creates a synergistic system for missing‑data estimation. The approach delivers statistically significant accuracy gains (averaging a 13 % improvement across models) and reduces the computational burden of the optimization phase. This hybrid methodology offers a practical, scalable solution for real‑world datasets where missing values are pervasive and accurate imputation is critical for downstream analysis.
Comments & Academic Discussion
Loading comments...
Leave a Comment