Exact Statistical Mechanical Investigation of a Finite Model Protein in its environment: A Small System Paradigm
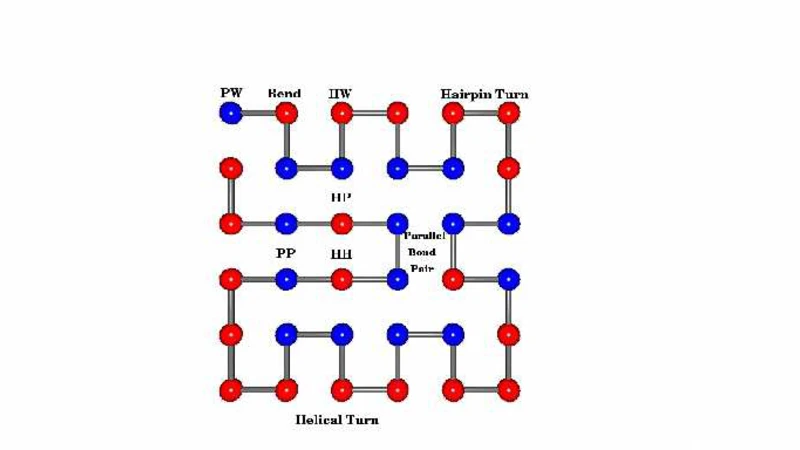
We consider a general incompressible finite model protein of size M in its environment, which we represent by a semiflexible copolymer consisting of amino acid residues classified into only two species (H and P, see text) following Lau and Dill. We allow various interactions between chemically unbonded residues in a given sequence and the solvent (water), and exactly enumerate the number of conformations W(E) as a function of the energy E on an infinite lattice under two different conditions: (i) we allow conformations that are restricted to be compact (known as Hamilton walk conformations), and (ii) we allow unrestricted conformations that can also be non-compact. It is easily demonstrated using plausible arguments that our model does not possess any energy gap even though it is supposed to exhibit a sharp folding transition in the thermodynamic limit. The enumeration allows us to investigate exactly the effects of energetics on the native state(s), and the effect of small size on protein thermodynamics and, in particular, on the differences between the microcanonical and canonical ensembles. We find that the canonical entropy is much larger than the microcanonical entropy for finite systems. We investigate the property of self-averaging and conclude that small proteins do not self-average. We also present results that (i) provide some understanding of the energy landscape, and (ii) shed light on the free energy landscape at different temperatures.
💡 Research Summary
The paper presents an exact statistical‑mechanical study of a finite lattice protein model that incorporates only two residue types, hydrophobic (H) and polar (P), following the classic Lau‑Dill HP framework. The authors treat the protein as an incompressible semiflexible copolymer of length M placed on an infinite lattice and enumerate every possible conformation, thereby obtaining the exact density of states W(E) for each energy E. Two distinct ensembles of conformations are considered. In the first, only compact configurations are allowed; mathematically these are Hamiltonian walks that visit every lattice site exactly once, representing the traditional “compact‑folded” assumption. In the second, the walk is unrestricted, permitting non‑compact, extended structures that more faithfully mimic the variety of real protein conformations in solution.
For each case the authors compute W(E) by exhaustive enumeration, which enables a direct comparison between microcanonical quantities (entropy S_micro(E)=k_B ln W(E)) and canonical quantities (free energy F(T)=−k_B T ln Z, entropy S_can(T)=−∂F/∂T). The analysis yields several key insights. First, the energy spectrum shows no gap: the density of states is continuous across the entire range of energies, contradicting the common picture of a two‑state folding transition that relies on an energy gap between native and unfolded ensembles. Second, for finite M the canonical entropy is substantially larger than the microcanonical entropy, reflecting the well‑known ensemble inequivalence in small systems. This difference grows roughly linearly with M, indicating that only in the thermodynamic limit do the two ensembles converge.
Third, the authors examine self‑averaging by sampling many random H/P sequences. The variance of thermodynamic observables (average energy, entropy) does not diminish with increasing M, demonstrating that small proteins do not self‑average; each sequence retains distinct thermodynamic signatures. This result provides a statistical‑mechanical justification for the empirical observation that protein function and stability are highly sequence‑specific.
Fourth, the exact density of states permits a detailed reconstruction of the energy landscape. In the compact (Hamiltonian‑walk) ensemble, low‑energy states cluster into a few deep minima, suggesting a funnel‑like free‑energy landscape that guides folding toward a unique or few native structures. By contrast, the unrestricted ensemble exhibits a broader band of low‑energy states, implying multiple competing minima and a more rugged landscape. Temperature‑dependent free‑energy profiles reveal sharp cross‑overs that resemble macroscopic folding transitions, even though the underlying system is finite.
Finally, the paper quantifies the entropy gap ΔS = S_can − S_micro and shows that it scales with system size, reinforcing the notion that finite‑size effects dominate protein thermodynamics at biologically relevant lengths. The authors conclude that exact enumeration, while computationally intensive, offers unparalleled insight into the interplay between sequence, energetics, and conformational statistics. Their findings challenge the applicability of thermodynamic limit arguments to real proteins, highlight the importance of ensemble choice for small systems, and provide a benchmark for future coarse‑grained or Monte‑Carlo studies of protein folding.
Comments & Academic Discussion
Loading comments...
Leave a Comment