Pseudoknot RNA Structures with Arc-Length $ge 3$
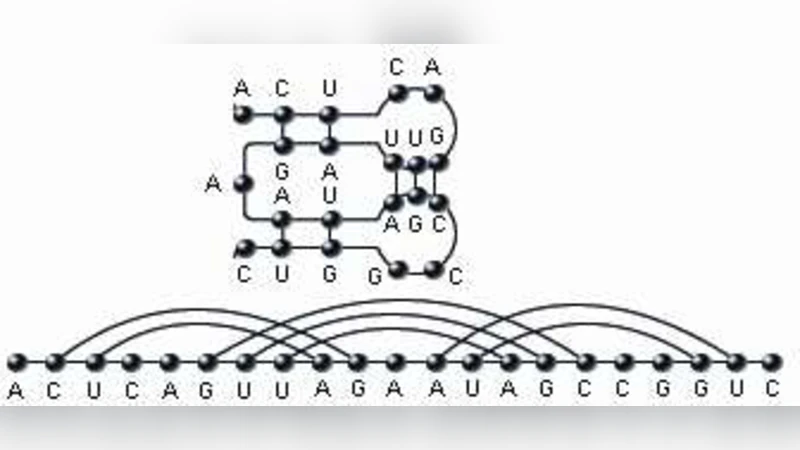
In this paper we study $k$-noncrossing RNA structures with arc-length $\ge 3$, i.e. RNA molecules in which for any $i$, the nucleotides labeled $i$ and $i+j$ ($j=1,2$) cannot form a bond and in which there are at most $k-1$ mutually crossing arcs. Let ${\sf S}{k,3}(n)$ denote their number. Based on a novel functional equation for the generating function $\sum{n\ge 0}{\sf S}{k,3}(n)z^n$, we derive for arbitrary $k\ge 3$ exponential growth factors and for $k=3$ the subexponential factor. Our main result is the derivation of the formula ${\sf S}{3,3}(n) \sim \frac{6.11170\cdot 4!}{n(n-1)…(n-4)} 4.54920^n$.
💡 Research Summary
The paper investigates a refined class of RNA secondary structures called k‑noncrossing RNA pseudoknot structures with arc‑length at least three. In this model, nucleotides at positions i and i + j may form a base pair only if j ≥ 3, thereby excluding the short arcs of length one and two that are biologically unrealistic for many RNAs. Additionally, the k‑noncrossing condition limits the diagram to contain at most k − 1 mutually crossing arcs, which captures the notion of limited pseudoknot complexity.
The authors denote by ({\sf S}_{k,3}(n)) the number of such structures on a sequence of length n and study the ordinary generating function
\
Comments & Academic Discussion
Loading comments...
Leave a Comment