A probabilistic regulatory network for the human immune system
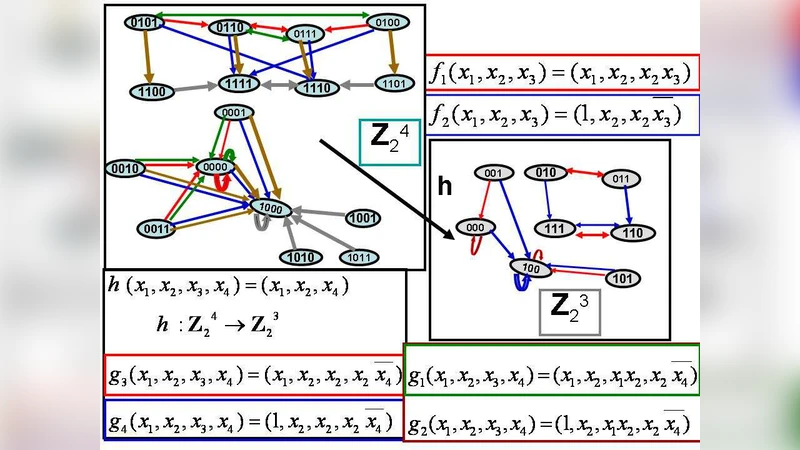
In this paper we made a review of some papers about probabilistic regulatory networks (PRN), in particular we introduce our concept of homomorphisms of PRN with an example of projection of a regulatory network to a smaller one. We apply the model PRN (or Probabilistic Boolean Network) to the immune system, the PRN works with two functions. The model called ““The B/T-cells interaction”” is Boolean, so we are really working with a Probabilistic Boolean Network. Using Markov Chains we determine the state of equilibrium of the immune response.
💡 Research Summary
The paper provides a two‑part contribution to the study of probabilistic regulatory networks (PRNs) and their application to immunology. In the first part the authors review the existing literature on PRNs, which are models that combine Boolean update functions with probabilistic selection to capture the stochastic nature of gene regulation, signaling pathways, and other cellular processes. They identify a key limitation of current approaches: as the number of genes or regulatory elements grows, the state space expands exponentially, making exact analysis intractable. To address this, the authors introduce the concept of a homomorphism between PRNs. A homomorphism is a mapping from the state space of a larger network to that of a smaller one that preserves the transition structure and the associated probabilities (or at least their aggregate effect). In mathematical terms, the transition matrix of the original network is transformed into a reduced matrix that retains the eigen‑spectral properties relevant for long‑term behavior. The paper illustrates the idea with a concrete example: a four‑gene network governed by two Boolean functions is projected onto a two‑gene network. The authors show how the probability weights of the original functions are redistributed in the reduced model and prove that the steady‑state distribution of the Markov chain is invariant under the homomorphism.
The second part applies this theoretical framework to a simplified model of the human immune response, specifically the interaction between B‑cells and T‑cells. The authors represent the system with two Boolean variables—B‑cell activation and T‑cell activation—and two regulatory functions that capture the mutual stimulation and inhibition observed in real immunological processes. Each function is assigned a selection probability (for example, 0.6 for one rule and 0.4 for the alternative), reflecting biological variability. By constructing the corresponding transition probability matrix, they treat the system as a discrete‑time Markov chain. Solving for the stationary distribution yields the long‑run probabilities of the four possible states: both cells inactive, only B‑cell active, only T‑cell active, and both cells active. The results indicate under which probabilistic regimes the immune response tends toward sustained activation (high probability of the both‑active state) versus quiescence or regulatory suppression (high probability of the both‑inactive state). This quantitative description of equilibrium provides insight into how stochastic regulatory choices can shape the overall immune dynamics.
Beyond the specific B/T‑cell example, the authors argue that homomorphism‑based reduction can be systematically applied to larger immunological networks that include cytokines, helper and regulatory T‑cells, and additional signaling pathways. By preserving the essential stochastic dynamics while drastically shrinking the state space, the approach makes exact Markov‑chain analysis feasible for systems that would otherwise require costly simulation or approximation methods.
The paper acknowledges several limitations. The Boolean abstraction reduces the rich, continuous range of signaling intensities to binary on/off states, potentially overlooking graded responses and multi‑step activation cascades. Moreover, the choice of transition probabilities is based on assumed values rather than experimentally calibrated data, which may affect the biological realism of the predictions. The authors suggest future work should incorporate multi‑valued logic, expand the repertoire of regulatory functions, and validate the models against time‑course immunological measurements.
In summary, the study makes three principal contributions: (1) it formalizes PRN homomorphisms and demonstrates their mathematical properties; (2) it provides a concrete method for reducing complex stochastic networks without losing essential dynamical information; and (3) it applies the reduced PRN framework to a Boolean model of B‑cell/T‑cell interaction, using Markov‑chain analysis to elucidate the equilibrium behavior of the immune response. This integration of network reduction theory with immunological modeling opens a promising avenue for scalable, quantitative analysis of large‑scale biological regulatory systems.