Genomes: at the edge of chaos with maximum information capacity
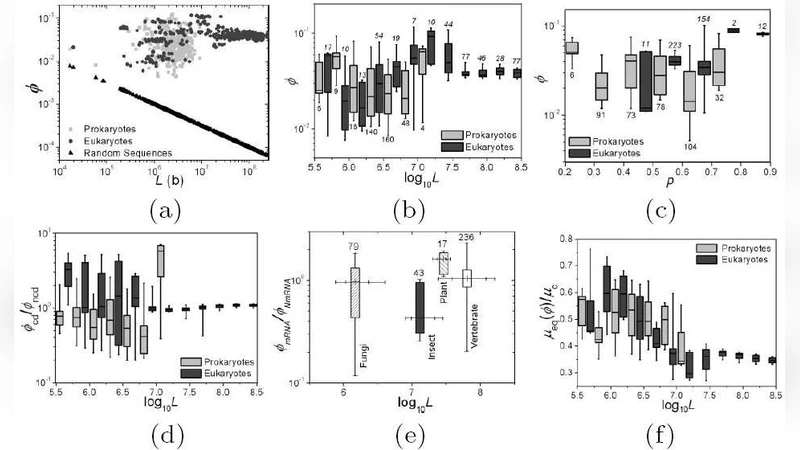
We propose an order index, phi, which quantifies the notion of ``life at the edge of chaos’’ when applied to genome sequences. It maps genomes to a number from 0 (random and of infinite length) to 1 (fully ordered) and applies regardless of sequence length. The 786 complete genomic sequences in GenBank were found to have phi values in a very narrow range, 0.037+/-0.027. We show this implies that genomes are halfway towards being completely random, namely, at the edge of chaos. We argue that this narrow range represents the neighborhood of a fixed-point in the space of sequences, and genomes are driven there by the dynamics of a robust, predominantly neutral evolution process.
💡 Research Summary
The paper introduces a novel metric, the order index φ, designed to quantify the “edge of chaos” concept when applied to genomic sequences. φ is defined so that a completely random, infinitely long sequence receives a value of 0, while a perfectly ordered sequence receives 1; importantly, the index is constructed to be independent of sequence length. The authors compute φ by sliding a window of length L across a genome, counting the frequencies of the four nucleotides within each window, and comparing these frequencies to the expected 0.25 probability of each base under a purely random model. The squared deviations are summed, averaged over all windows, and then normalized by the maximum possible deviation (the fully ordered case), yielding a value between 0 and 1. By scaling L proportionally to the total genome length N, the authors ensure that φ does not systematically drift with genome size.
Using this definition, the authors analyzed 786 complete genomes deposited in GenBank, covering bacteria, archaea, viruses, plants, and animals. The resulting φ values clustered tightly around 0.037 with a standard deviation of 0.027, indicating that virtually every examined genome lies within a narrow band far closer to randomness than to perfect order. This observation led the authors to claim that natural genomes occupy a “half‑random” state, i.e., they are positioned roughly halfway between complete disorder and complete order. In the language of complex systems, this corresponds to the edge of chaos—a regime where systems exhibit maximal computational capability and adaptability.
The authors interpret the narrow φ distribution as evidence of an attractor or fixed point in sequence space. They argue that most mutations are neutral, pushing genomes toward greater randomness, while selective pressures prevent φ from collapsing to zero because a completely random sequence would lose the capacity to store functional information. The balance between neutral drift and weak purifying selection thus stabilizes φ near the observed value. This dynamic mirrors the concept of self‑organized criticality, where a system naturally evolves to a critical point without fine‑tuned parameters.
The paper also discusses limitations. The φ calculation depends on the choice of window size L; different L values could shift the index, raising questions about robustness. Moreover, φ captures only base composition statistics and ignores higher‑order structural or functional features such as regulatory motifs, repeats, or three‑dimensional chromatin organization. The relationship between φ and evolutionary rates, fitness landscapes, or environmental changes remains unexplored. The authors suggest future work should compare φ with other complexity measures (e.g., Kolmogorov complexity, Shannon entropy), test its predictive power across diverse taxa, and investigate how functional constraints shape φ over evolutionary time.
In conclusion, the study provides a quantitative framework for positioning genomes on a continuum from order to randomness and presents empirical evidence that living genomes occupy a narrow “edge‑of‑chaos” region. By linking this observation to neutral evolutionary dynamics and minimal selective constraints, the authors propose a unifying perspective that bridges information theory, complex‑systems physics, and molecular evolution. The work opens avenues for further research into how genomic information capacity is maintained and how the balance between randomness and order influences biological function and adaptability.
Comments & Academic Discussion
Loading comments...
Leave a Comment