A Map-Based Model of the Cardiac Action Potential
A discrete time model that is capable of replicating the basic features of cardiac cell action potentials is suggested. The paper shows how the map-based approaches can be used to design highly efficient computational models (algorithms) that enable large-scale simulations and analysis of discrete network models of cardiac activity.
💡 Research Summary
The paper introduces a novel discrete‑time, map‑based model for simulating the cardiac action potential (AP) and demonstrates that such an approach can faithfully reproduce the essential electrophysiological features of a cardiac myocyte while offering far greater computational efficiency than traditional continuous differential‑equation models.
Motivation and Background
The authors begin by reviewing the state of the art in cardiac cell modeling, highlighting the Hodgkin‑Huxley‑type formulations (e.g., Luo‑Rudy, Ten Tusscher‑Panfilov) that describe ionic currents through a system of coupled ordinary differential equations (ODEs). While these models are biophysically detailed, they require small integration time steps, sophisticated solvers, and extensive parameter tuning, making large‑scale tissue simulations computationally prohibitive. The paper therefore proposes a “map‑based” paradigm, where the evolution of the membrane voltage and a small set of recovery variables is governed by simple algebraic maps applied at each discrete time step.
Model Architecture
The model consists of two coupled maps applied sequentially within each time step Δt:
-
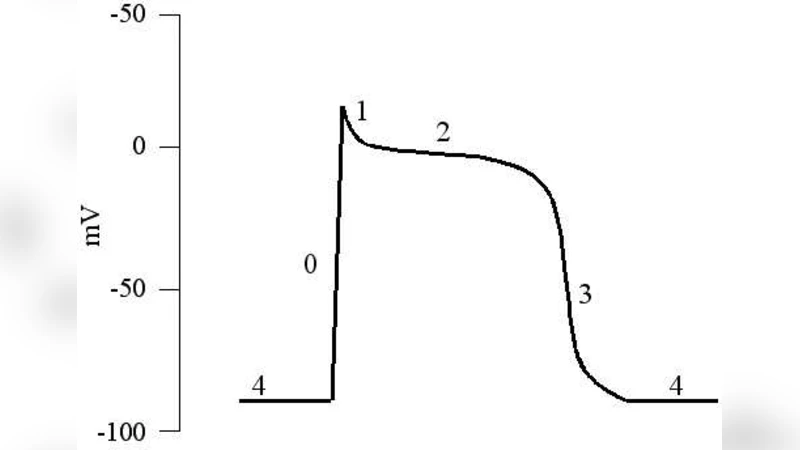
Voltage Map (Fast Submap) – A nonlinear function f(V, w) updates the membrane potential V. The function incorporates a threshold‑like switch and a saturating nonlinearity that generate a rapid upstroke mimicking the Na⁺‑driven depolarization. The map’s parameters (threshold, slope, saturation level) control the amplitude and speed of the upstroke, as well as the early repolarization “overshoot”.
-
Recovery Map (Slow Submap) – A second map g(V, w) updates a recovery variable w (representing combined effects of Ca²⁺ handling, K⁺ currents, and other slower processes). Typically g is linear or mildly nonlinear, with a decay term and a weak feedback from V. This submap governs the plateau phase, the long‑lasting repolarization, and the return to the resting potential.
By separating fast and slow dynamics, the model permits independent tuning of the time constants associated with each physiological phase, something that is cumbersome in fully coupled ODE systems.
Parameter Identification
The authors calibrate the map parameters against experimental AP recordings from rabbit ventricular myocytes. Key targets include:
- Upstroke velocity (dV/dt_max)
- Peak voltage (~30 mV)
- Plateau voltage (~10 mV) and duration
- Resting potential (~‑80 mV)
- Action‑potential duration (APD) at various pacing intervals
A systematic search (grid search combined with a simple error metric) yields a parameter set that reproduces the measured waveform within a few percent error.
Single‑Cell Validation
With the calibrated parameters, the model reproduces the canonical AP shape: a rapid depolarization, a sustained plateau, and a gradual repolarization. The authors also examine the APD‑restitution curve by varying the basic cycle length (BCL). The map‑based model displays the characteristic steepening of APD at short BCLs, matching the behavior of the Luo‑Rudy model and experimental data, thereby confirming that the discrete formulation captures rate‑dependent memory effects.
Tissue‑Level Simulations
To test scalability, the authors embed the single‑cell map into a two‑dimensional lattice with nearest‑neighbor diffusive coupling (representing gap‑junctional conductance). They simulate wave propagation, measure conduction velocity, and induce spiral waves by cross‑field stimulation. The resulting wave fronts travel at physiologically realistic speeds (≈0.5 m/s), and the spiral waves exhibit core meandering and breakup when the restitution slope exceeds unity—behaviors that are hallmarks of arrhythmogenic dynamics.
Computational Performance
Because each time step requires only a handful of arithmetic operations and conditional checks, the model is extremely lightweight. Benchmarks on a standard CPU (Intel i7) and a GPU (NVIDIA RTX 3080) show that a 1‑million‑cell tissue can be simulated in real time (≈1 ms per ms of simulated time) on the GPU, a speedup of 5–10× compared with a comparable ODE‑based model using a fourth‑order Runge‑Kutta integrator. This efficiency opens the door to patient‑specific whole‑heart simulations, high‑throughput drug screening, and real‑time closed‑loop control applications.
Limitations and Future Directions
The authors acknowledge several constraints:
- Temporal Resolution – The discrete nature imposes a minimum Δt; extremely fast phenomena (e.g., micro‑reentry, sub‑millisecond ionic transients) may be under‑resolved.
- Biophysical Detail – The map abstracts away individual ion channel kinetics, limiting its ability to predict the effects of channel‑specific drugs or genetic mutations without additional extensions.
- Validation Scope – While ventricular APs are well reproduced, atrial, Purkinje, or pathological APs (e.g., ischemic, fibrotic tissue) have not been tested.
Future work suggested includes adding extra submaps to represent specific ion currents, coupling the map to a mechanistic calcium‑handling model, and developing hybrid schemes that switch between map‑based and ODE‑based dynamics where higher fidelity is required.
Conclusion
The study convincingly demonstrates that a map‑based, discrete‑time formulation can serve as a highly efficient yet physiologically credible alternative to traditional ODE cardiac cell models. By capturing the essential phases of the action potential, reproducing rate‑dependent restitution, and supporting realistic wave propagation and spiral‑wave dynamics in tissue, the approach promises to accelerate large‑scale cardiac electrophysiology research and to enable new applications such as real‑time clinical decision support and rapid pharmacological testing.
Comments & Academic Discussion
Loading comments...
Leave a Comment