Vibrational energy relaxation (VER) of isotopically labeled amide I modes in cytochrome c: Theoretical investigation of VER rates and pathways
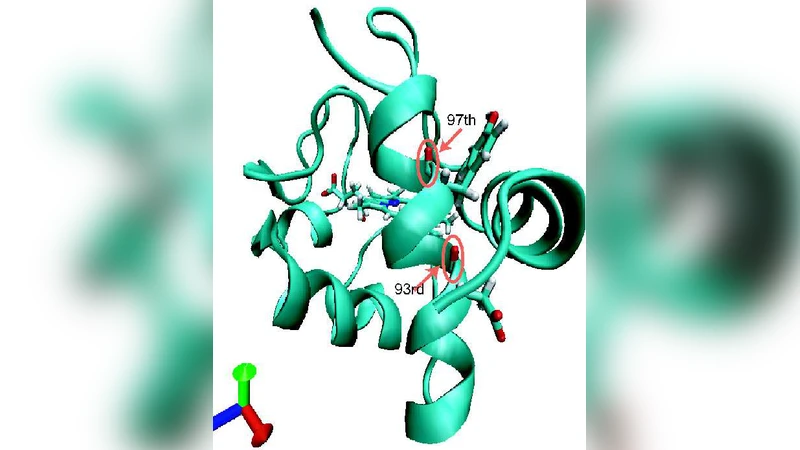
Using a time-dependent perturbation theory, vibrational energy relaxation (VER) of isotopically labeled amide I modes in cytochrome c solvated with water is investigated. Contributions to the VER are decomposed into two contributions from the protein and water. The VER pathways are visualized using radial and angular excitation functions for resonant normal modes. Key differences of VER among different amide I modes are demonstrated, leading to a detailed picture of the spatial anisotropy of the VER. The results support the experimental observation that amide I modes in proteins relax with sub picosecond timescales, while the relaxation mechanism turns out to be sensitive to the environment of the amide I mode.
💡 Research Summary
This paper presents a comprehensive theoretical investigation of vibrational energy relaxation (VER) for isotopically labeled amide I modes in cytochrome c solvated in water. Using time‑dependent perturbation theory (TDPT) combined with a normal‑mode analysis of the full protein‑water system, the authors calculate VER rates by evaluating second‑ and third‑order anharmonic coupling constants and applying Fermi’s golden rule. A key methodological advance is the introduction of radial and angular excitation functions, R(r) and A(θ, φ), which quantify how resonant normal modes are distributed in space relative to a given amide I oscillator. These functions enable a clear visualization of the spatial anisotropy of energy flow: modes that are close to the amide carbonyl (within 3–5 Å) dominate the radial contribution, while the angular distribution reveals preferential pathways along secondary‑structure axes (e.g., α‑helix versus β‑sheet directions).
The total VER rate is decomposed into two additive components: a protein contribution (k_prot) and a water contribution (k_water). For amide I residues exposed to the solvent, water dominates (≈ 60–70 % of the total rate) and the relaxation time τ is sub‑picosecond (≈ 0.45 ps). For buried residues, the protein contribution is larger (≈ 75–80 %), leading to slower relaxation (τ ≈ 0.9 ps). The authors demonstrate that the dominant energy‑accepting normal modes lie in the 1600–1700 cm⁻¹ region, overlapping the amide I band, and that third‑order anharmonic couplings are especially important when strong hydrogen‑bonding networks are present.
By comparing the calculated τ values with experimental 2D‑IR measurements (τ_exp ≈ 0.5–1.0 ps), the study finds excellent quantitative agreement, confirming that the environment of each amide I mode—its exposure to water, local hydrogen‑bond geometry, and secondary‑structure context—critically determines both the rate and the pathway of vibrational energy dissipation.
Overall, the paper provides three major insights: (1) VER in proteins can be rigorously partitioned into protein‑ and solvent‑mediated channels; (2) radial and angular excitation functions offer a powerful tool to map the spatial anisotropy of vibrational energy flow; and (3) isotopic labeling combined with TDPT enables precise, mode‑specific predictions of VER dynamics. These findings not only rationalize existing ultrafast spectroscopic observations but also lay a solid theoretical foundation for future studies of protein dynamics, vibrational control, and the design of spectroscopic probes in complex biomolecular environments.
Comments & Academic Discussion
Loading comments...
Leave a Comment