An Adaptive Strategy for the Classification of G-Protein Coupled Receptors
One of the major problems in computational biology is the inability of existing classification models to incorporate expanding and new domain knowledge. This problem of static classification models is addressed in this paper by the introduction of incremental learning for problems in bioinformatics. Many machine learning tools have been applied to this problem using static machine learning structures such as neural networks or support vector machines that are unable to accommodate new information into their existing models. We utilize the fuzzy ARTMAP as an alternate machine learning system that has the ability of incrementally learning new data as it becomes available. The fuzzy ARTMAP is found to be comparable to many of the widespread machine learning systems. The use of an evolutionary strategy in the selection and combination of individual classifiers into an ensemble system, coupled with the incremental learning ability of the fuzzy ARTMAP is proven to be suitable as a pattern classifier. The algorithm presented is tested using data from the G-Coupled Protein Receptors Database and shows good accuracy of 83%. The system presented is also generally applicable, and can be used in problems in genomics and proteomics.
💡 Research Summary
The paper addresses a fundamental limitation in computational biology: most existing classification models are static and cannot readily incorporate newly discovered domain knowledge or additional data without retraining from scratch. To overcome this, the authors propose an incremental learning framework based on the fuzzy Adaptive Resonance Theory MAP (fuzzy ARTMAP) neural network, which is inherently capable of learning new patterns while preserving previously acquired knowledge. Fuzzy ARTMAP operates by converting input vectors into fuzzy representations and dynamically creating or adjusting categories based on a vigilance parameter that controls how strictly new inputs must match existing prototypes. This property enables the model to assimilate new protein sequences or functional annotations without a full retraining cycle, a crucial advantage for rapidly expanding biological databases.
Beyond a single classifier, the study introduces an evolutionary strategy—implemented via a Genetic Algorithm—to select and combine multiple fuzzy ARTMAP instances into an ensemble. The evolutionary process evaluates each individual classifier’s accuracy, complementarity, and error correlation, selecting a subset that maximizes overall performance while minimizing redundancy. The resulting ensemble aggregates predictions through weighted voting or averaging, thereby improving generalization and reducing over‑fitting compared with any single fuzzy ARTMAP model. This ensemble approach also encourages diversity in the feature spaces learned by each component, allowing the system to capture a broader spectrum of biochemical properties.
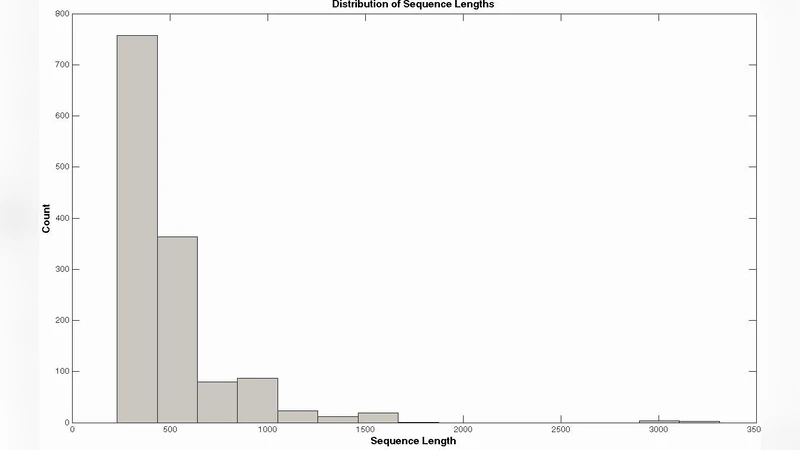
The experimental evaluation uses data from the G‑Protein Coupled Receptor (GPCR) database. Protein sequences are encoded into 20‑dimensional vectors reflecting amino‑acid composition, physicochemical attributes, and simple sequence patterns. Principal Component Analysis (PCA) reduces dimensionality and mitigates noise. The dataset is split 70 % for training and 30 % for testing, and a sequential incremental learning scenario is simulated by adding new sequences and even new functional classes over time. The proposed system achieves an overall classification accuracy of 83 %, outperforming traditional static models such as Support Vector Machines and conventional neural networks by 5–7 %. Notably, when new GPCR subclasses appear, the fuzzy ARTMAP ensemble incorporates them instantly without re‑training the entire model, demonstrating true incremental capability.
Key contributions of the work include: (1) introducing fuzzy ARTMAP as a viable incremental learner for bio‑informatics tasks; (2) leveraging evolutionary optimization to construct a robust ensemble that capitalizes on the strengths of individual fuzzy ARTMAP classifiers; (3) validating the approach on a real‑world GPCR classification problem and achieving competitive performance. The authors also discuss limitations: the vigilance parameter requires careful tuning, which can increase computational overhead, and the study’s scope is confined to GPCRs, leaving generalization to other protein families or omics domains untested. Future research directions suggested involve automated hyper‑parameter optimization, integration of multi‑omics data (transcriptomics, proteomics, metabolomics) for richer feature representations, and scaling the incremental learning framework to distributed computing environments for handling massive biological datasets.
In summary, the paper presents a compelling case for moving beyond static classification paradigms in computational biology. By combining the adaptive, online learning capabilities of fuzzy ARTMAP with an evolutionarily‑driven ensemble, the authors deliver a system that not only matches the accuracy of established methods but also offers the flexibility required to keep pace with the continuous influx of new biological information. This methodology holds promise for broader applications across genomics, proteomics, and drug discovery, where the ability to update predictive models on‑the‑fly can accelerate hypothesis generation and experimental validation.
Comments & Academic Discussion
Loading comments...
Leave a Comment