Kinetics of cell division in epidermal maintenance
The rules governing cell division and differentiation are central to understanding the mechanisms of development, aging and cancer. By utilising inducible genetic labelling, recent studies have shown that the clonal population in transgenic mouse epidermis can be tracked in vivo. Drawing on these results, we explain how clonal fate data may be used to infer the rules of cell division and differentiation underlying the maintenance of adult murine tail-skin. We show that the rates of cell division and differentiation may be evaluated by considering the long-time and short-time clone fate data, and that the data is consistent with cells dividing independently rather than synchronously. Motivated by these findings, we consider a mechanism for cancer onset based closely on the model for normal adult skin. By analysing the expected changes to clonal fate in cancer emerging from a simple two-stage mutation, we propose that clonal fate data may provide a novel method for studying the earliest stages of the disease.
💡 Research Summary
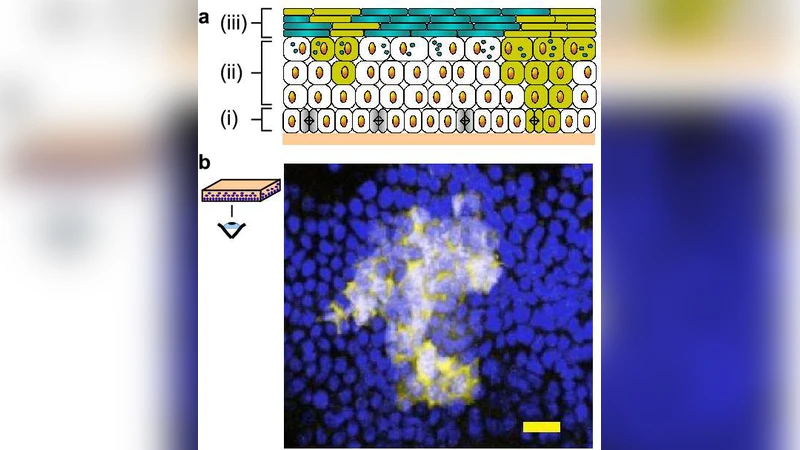
The paper addresses a fundamental question in tissue biology: how are cell division and differentiation coordinated to maintain the adult epidermis, and how might perturbations of this balance give rise to cancer? Using an inducible Cre‑ER system in transgenic mice (K14‑CreER;Rosa26‑YFP), the authors permanently label a sparse subset of basal keratinocytes and then follow the fate of each labeled clone over periods ranging from one day to several weeks. By counting the number of YFP‑positive cells that remain in the basal layer versus those that have migrated suprabasally, they obtain a time‑resolved distribution of clone sizes and compositions.
To extract quantitative rules from these data, the authors formulate a stochastic birth‑death (or “division‑differentiation”) model. In this framework each basal cell independently undergoes symmetric division at a rate λ, and with probability proportional to a second rate γ it differentiates and leaves the basal compartment. The model predicts two observable signatures: (i) the short‑time distribution of clone sizes, which is dominated by the ratio λ/γ, and (ii) the long‑time behavior, characterized by a constant clone extinction probability and a linear increase of the mean clone size with time. Maximum‑likelihood fitting of the experimental data to the analytical solutions of the model yields λ≈0.12 day⁻¹ and γ≈0.08 day⁻¹, indicating that basal cells divide roughly once every eight days and differentiate at a comparable pace. Importantly, the fit is markedly better for the independent‑division hypothesis than for models that assume synchronized or coordinated bursts of division, confirming that epidermal homeostasis is achieved through stochastic, cell‑autonomous events.
Having established a robust quantitative baseline for normal skin, the authors then explore how early oncogenic mutations would alter clonal dynamics. They introduce a minimal two‑stage mutation scheme: the first hit disables a growth‑suppressive pathway (e.g., p53 loss), and the second activates a proliferative driver (e.g., Ras activation). Cells that acquire both hits are assigned altered parameters λ′>λ and γ′<γ, reflecting faster division and slower differentiation. Stochastic simulations of this “cancer‑seed” model reveal a characteristic shift in the clone‑size distribution: while early mutant clones are indistinguishable from wild‑type ones, after a critical time point a heavy tail emerges, representing a subpopulation of rapidly expanding clones that persist far longer than normal clones. The authors argue that this tail, detectable by routine clonal fate analysis, could serve as a sensitive early‑detection marker for incipient skin cancers, long before histopathological changes become apparent.
The study therefore makes three major contributions. First, it demonstrates that high‑resolution clonal tracing combined with simple birth‑death mathematics can yield precise estimates of division and differentiation rates in a living tissue. Second, it provides strong experimental support for the notion that epidermal maintenance is governed by independent, stochastic cell behavior rather than coordinated cycles. Third, it translates the quantitative framework into a predictive tool for cancer biology, showing that subtle alterations in the underlying kinetic parameters produce measurable changes in clone fate that could be exploited for early diagnosis or for testing the impact of preventive interventions. By bridging experimental lineage tracing, statistical physics, and oncogenic modeling, the paper sets a new standard for dissecting tissue dynamics and for leveraging those insights to confront disease at its earliest stages.
Comments & Academic Discussion
Loading comments...
Leave a Comment