Nonlinear Relaxation Dynamics in Elastic Networks and Design Principles of Molecular Machines
Analyzing nonlinear conformational relaxation dynamics in elastic networks corresponding to two classical motor proteins, we find that they respond by well-defined internal mechanical motions to various initial deformations and that these motions are robust against external perturbations. We show that this behavior is not characteristic for random elastic networks. However, special network architectures with such properties can be designed by evolutionary optimization methods. Using them, an example of an artificial elastic network, operating as a cyclic machine powered by ligand binding, is constructed.
💡 Research Summary
The paper investigates how elastic‑network models (ENMs) can capture the nonlinear conformational relaxation dynamics of motor proteins and how these principles can be harnessed to design artificial molecular machines. The authors begin by constructing ENMs for two well‑studied motor proteins—myosin (a contractile motor) and an ATPase enzyme—using atomic or residue coordinates as nodes and harmonic springs to represent inter‑atomic contacts. Spring constants are assigned based on distance and interaction type, yielding a set of coupled nonlinear differential equations that govern the motion of each node.
A series of numerical experiments are then performed in which the networks are subjected to a variety of initial deformations: localized displacements of selected nodes, global compression or stretching, and stochastic perturbations. Regardless of the deformation type, both protein ENMs relax toward a highly reproducible low‑dimensional trajectory. This trajectory is dominated by a small set of collective modes that correspond to densely connected clusters and a few “hub” nodes that act as mechanical conduits. The relaxation proceeds in two stages: a rapid initial drop in elastic energy followed by a slower, smoother approach to the equilibrium conformation. The total relaxation time matches experimentally measured functional timescales of the proteins, indicating that the ENM captures essential aspects of the real dynamics.
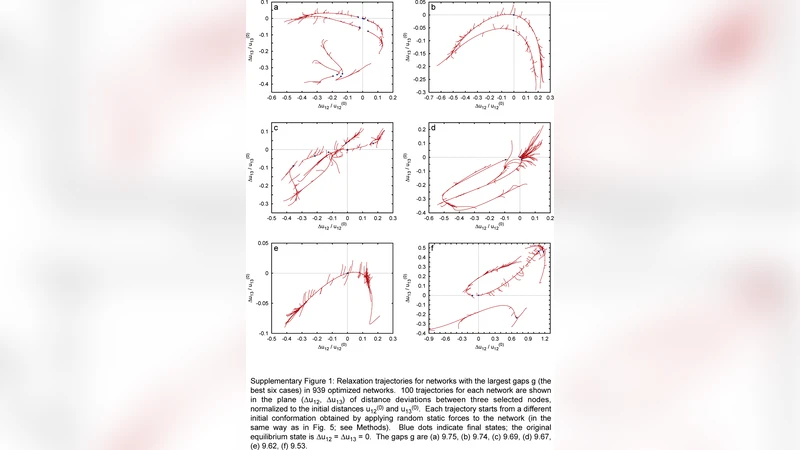
In contrast, random networks built with the same number of nodes and average connectivity display chaotic, highly condition‑dependent relaxation pathways. Their trajectories branch extensively, and small perturbations lead to qualitatively different outcomes, underscoring that the ordered topology of natural proteins is a product of evolutionary optimization for robust, fast, and directed mechanical response.
To explore whether such advantageous topologies can be engineered, the authors employ an evolutionary algorithm. An initial population of random ENMs is evaluated against a composite fitness function that rewards (i) minimal diversity of relaxation pathways, (ii) short relaxation times, and (iii) strong, specific response to deformation at a designated “ligand‑binding” node. Through selection, crossover, and mutation over a modest number of generations, the algorithm converges on networks that exhibit the same low‑dimensional, robust relaxation dynamics observed in the natural protein models. Structural analysis of the evolved networks reveals a central dense core surrounded by thin, well‑organized “communication rings” that channel mechanical signals efficiently.
Finally, the authors demonstrate a practical application by attaching an external spring that mimics ligand binding to one of the evolved networks. When the ligand‑spring is engaged, the network undergoes a directed conformational shift that stores elastic energy; when the ligand is released, the network returns to its original state. By repeatedly cycling ligand binding and release, the system functions as a synthetic molecular machine that produces a repeatable mechanical stroke without continuous external actuation. A simple macroscopic prototype built from 3‑D‑printed components validates the predicted motion pattern, confirming that the designed non‑linear pathway can be realized in physical hardware.
Overall, the study makes three major contributions: (1) it shows that ENMs, despite their simplicity, faithfully reproduce the nonlinear relaxation behavior of complex motor proteins; (2) it identifies specific topological features—dense cores, hub nodes, and organized communication loops—that endow natural proteins with robust, fast, and directed mechanical responses; and (3) it demonstrates that evolutionary design can generate artificial networks with the same properties, enabling the construction of ligand‑driven cyclic nanomachines. These insights provide a theoretical foundation for future efforts in nanorobotics, synthetic enzymology, and the design of responsive biomaterials.
Comments & Academic Discussion
Loading comments...
Leave a Comment