Ion transport through cell membrane channels
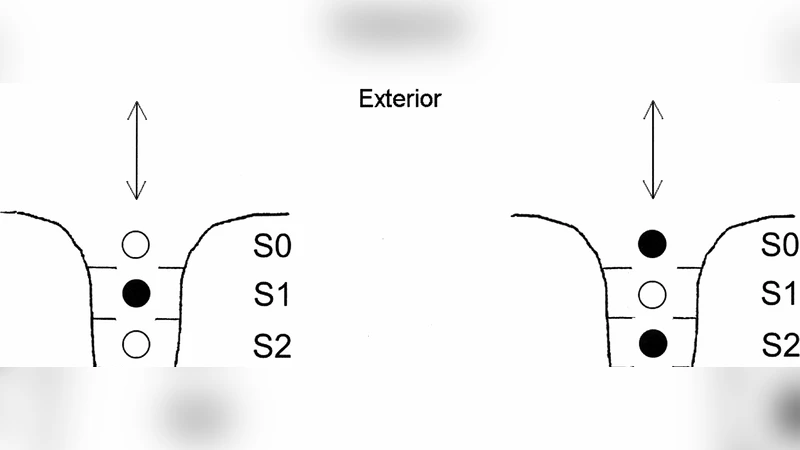
We discuss various models of ion transport through cell membrane channels. Recent experimental data shows that sizes of ion channels are compared to those of ions and that only few ions may be simultaneously in any single channel. Theoretical description of ion transport in such channels should therefore take into account interactions between ions and between ions and channel proteins. This is not satisfied by macroscopic continuum models based on Poisson-Nernst-Planck equations. More realistic descriptions of ion transport are offered by microscopic Brownian and molecular dynamics. One should also take into account a dynamical character of the channel structure. This is not yet addressed in the literature
💡 Research Summary
The paper provides a critical review of ion transport modeling in cell‑membrane channels, emphasizing that the dimensions of most biological channels are comparable to the size of the ions they conduct. Consequently, only one or a few ions can occupy a channel simultaneously, making ion‑ion and ion‑protein interactions essential to the transport process. Traditional macroscopic continuum approaches, particularly the Poisson‑Nernst‑Planck (PNP) equations, treat ions as independent particles moving in a continuous electric field and ignore these microscopic interactions. The authors demonstrate that PNP fails to reproduce key experimental observations such as single‑file movement, voltage‑dependent selectivity, and conductance blockades that arise from strong electrostatic repulsion, dehydration, and fixed charges within the narrow pore.
To overcome these limitations, the paper advocates for microscopic stochastic methods. Brownian dynamics (BD) simulations model each ion as a discrete particle, explicitly calculating Coulomb forces and the electric field derived from the Poisson equation. By incorporating realistic pore geometry and fixed charge distributions, BD can capture ion selectivity and voltage‑driven gating that PNP cannot. For an even more detailed description, molecular dynamics (MD) simulations are recommended. MD treats water molecules, channel protein atoms, and ions at the atomic level, allowing the system to evolve under realistic force fields. This approach reveals how water re‑orientation, protein side‑chain flexibility, and conformational changes of the channel gate influence ion permeation.
A major gap identified in the literature is the treatment of channel structure as static. Experimental evidence shows that channels undergo thermal fluctuations, voltage‑induced conformational shifts, and ligand‑binding–driven opening/closing. These dynamic changes provide feedback to the ion flux and must be incorporated into any predictive model. The authors argue for a multiscale framework that couples macroscopic electrostatic calculations with atomistic MD, possibly augmented by machine‑learning surrogates to accelerate long‑time simulations.
In conclusion, the paper calls for a paradigm shift from continuum to fully microscopic, dynamically coupled models. Such models are expected to improve our understanding of fundamental biophysical processes, guide the rational design of selective nanofluidic devices, and aid in the development of new pharmacological agents targeting ion channels.
Comments & Academic Discussion
Loading comments...
Leave a Comment