The Hypercube of Life: How Protein Stability Imposes Limits on Organism Complexity and Speed of Molecular Evolution
Classical population genetics a priori assigns fitness to alleles without considering molecular or functional properties of proteins that these alleles encode. Here we study population dynamics in a model where fitness can be inferred from physical properties of proteins under a physiological assumption that loss of stability of any protein encoded by an essential gene confers a lethal phenotype. Accumulation of mutations in organisms containing Gamma genes can then be represented as diffusion within the Gamma dimensional hypercube with adsorbing boundaries which are determined, in each dimension, by loss of a protein stability and, at higher stability, by lack of protein sequences. Solving the diffusion equation whose parameters are derived from the data on point mutations in proteins, we determine a universal distribution of protein stabilities, in agreement with existing data. The theory provides a fundamental relation between mutation rate, maximal genome size and thermodynamic response of proteins to point mutations. It establishes a universal speed limit on rate of molecular evolution by predicting that populations go extinct (via lethal mutagenesis) when mutation rate exceeds approximately 6 mutations per essential part of genome per replication for mesophilic organisms and 1 to 2 mutations per genome per replication for thermophilic ones. Further, our results suggest that in absence of error correction, modern RNA viruses and primordial genomes must necessarily be very short. Several RNA viruses function close to the evolutionary speed limit while error correction mechanisms used by DNA viruses and non-mutant strains of bacteria featuring various genome lengths and mutation rates have brought these organisms universally about 1000 fold below the natural speed limit.
💡 Research Summary
The paper challenges the conventional population‑genetics view that fitness is an abstract property assigned to alleles, and instead builds a mechanistic link between the thermodynamic stability of essential proteins and organismal fitness. The authors assume that loss of stability (ΔG above a critical threshold) in any protein encoded by an essential gene results in a lethal phenotype. Under this assumption, the accumulation of point mutations in a genome containing Γ essential genes can be represented as a random walk within a Γ‑dimensional hypercube, where each axis corresponds to the stability of one protein.
Two absorbing boundaries delimit the hypercube: (i) a lower‑stability boundary at which a protein becomes unstable enough to cause death, and (ii) an upper‑stability boundary imposed by the finite number of possible amino‑acid sequences. Mutations cause small displacements (ΔΔG) along each axis; the mean and variance of these displacements are taken from empirical measurements of single‑mutation effects on protein stability. Solving the diffusion equation
∂p/∂t = D∇²p – λp
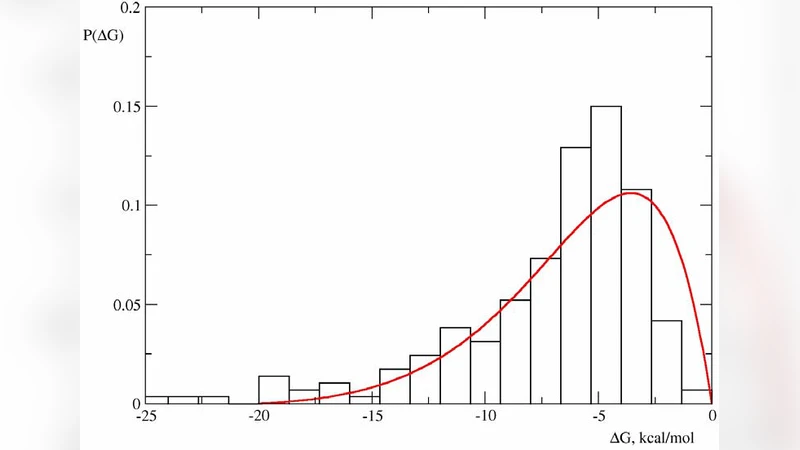
with the absorbing boundary conditions yields a stationary probability distribution of protein stabilities. The authors show that this distribution matches large‑scale experimental data (e.g., ProTherm, FoldX), thereby validating the model.
From the stationary solution they derive a universal relationship linking mutation rate (μ), the number of essential genes (Γ), and the average thermodynamic response of proteins to mutation (⟨ΔΔG⟩). The key result is a “lethal mutagenesis” threshold: when the expected loss of stability per replication reaches 1/Γ, the whole population collapses. Plugging typical values for mesophilic organisms (⟨ΔΔG⟩ ≈ 0.9 kcal mol⁻¹, μ ≈ 10⁻⁹ mut nt⁻¹ rep⁻¹) gives an upper bound of roughly six mutations per essential genome per replication. For thermophiles, whose proteins already sit near the stability ceiling, the threshold drops to 1–2 mutations per genome per replication. This defines a universal “speed limit” on molecular evolution.
The model has immediate biological implications. RNA viruses, which lack proofreading polymerases, typically have mutation rates of 10⁻⁴–10⁻³ mut nt⁻¹ rep⁻¹. Their genomes (a few thousand to tens of thousands of nucleotides) sit close to the derived limit, explaining why many RNA viruses evolve extremely rapidly and why they are vulnerable to lethal mutagenesis strategies (e.g., mutagenic nucleoside analogs). In contrast, DNA viruses and bacteria possess error‑correction mechanisms (DNA polymerase proofreading, viral exonucleases) that reduce μ to 10⁻⁶–10⁻⁸ mut nt⁻¹ rep⁻¹. Consequently, their actual evolutionary rates are about a thousand‑fold below the theoretical maximum, providing a safety margin that permits larger genomes and more complex lifestyles.
The authors also extrapolate to primordial life. In an RNA‑world scenario without any error‑correction, the lethal‑mutagenesis bound forces genomes to be only a few hundred nucleotides long, consistent with the size of ribozymes and the smallest extant RNA viruses. Thus, the model offers a quantitative framework linking protein physics, genome size, and evolutionary dynamics across the tree of life.
Beyond the specific findings, the study introduces a powerful statistical‑mechanics framework: a high‑dimensional diffusion process with absorbing boundaries. This approach can be extended to incorporate additional constraints such as metabolic network robustness, protein‑protein interaction networks, or environmental stressors (e.g., temperature shifts). By doing so, future work could derive analogous speed limits for other layers of cellular organization.
In summary, the paper provides (1) a physically grounded derivation of a universal distribution of protein stabilities, (2) a quantitative link between mutation rate, essential genome size, and protein thermodynamics, (3) a clear prediction of a mutation‑rate ceiling (≈6 mutations per essential genome for mesophiles, 1–2 for thermophiles), (4) an explanation for why RNA viruses operate near this ceiling while DNA‑based organisms stay far below it, and (5) a broader conceptual bridge between molecular biophysics and evolutionary theory, opening avenues for interdisciplinary research on the fundamental limits of biological complexity and adaptability.
Comments & Academic Discussion
Loading comments...
Leave a Comment