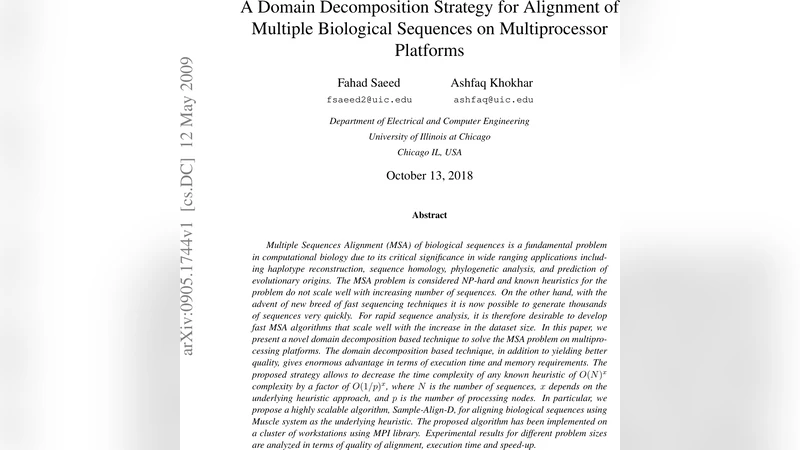
Q-Bio-Qm


Optimal metabolic pathway activation
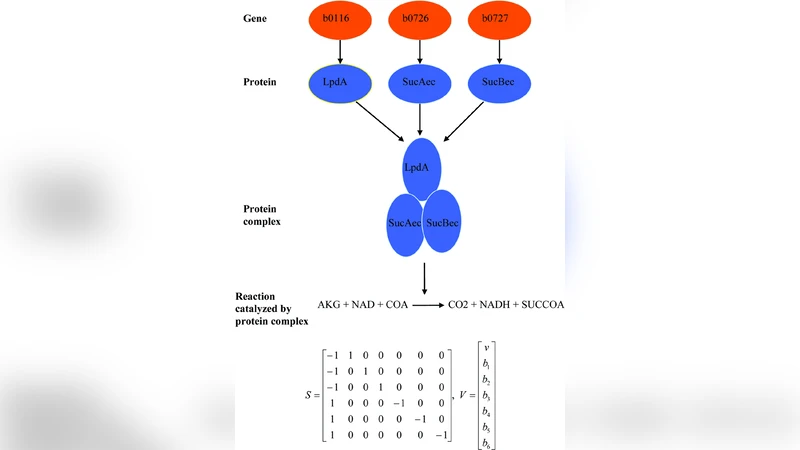
On the accessibility of adaptive phenotypes of a bacterial metabolic network

Estimation of biochemical network parameter distributions in cell populations

Computational Modeling for the Activation Cycle of G-proteins by G-protein-coupled Receptors
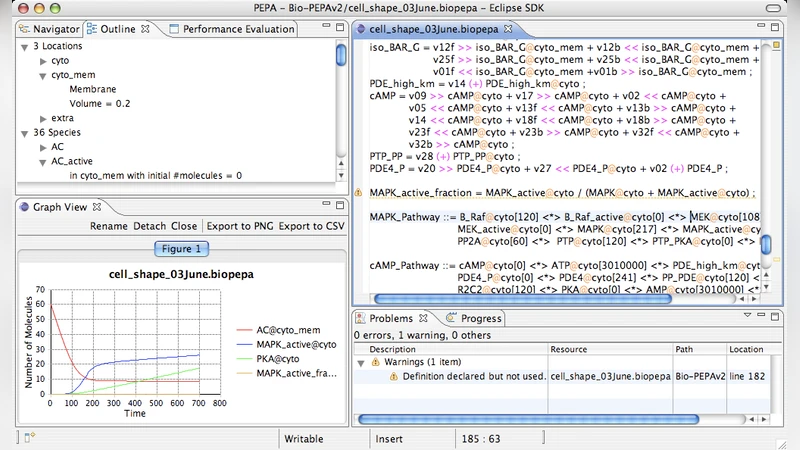
A compartmental model of the cAMP/PKA/MAPK pathway in Bio-PEPA

Coherent motion of stereocilia assures the concerted gating of hair-cell transduction channels

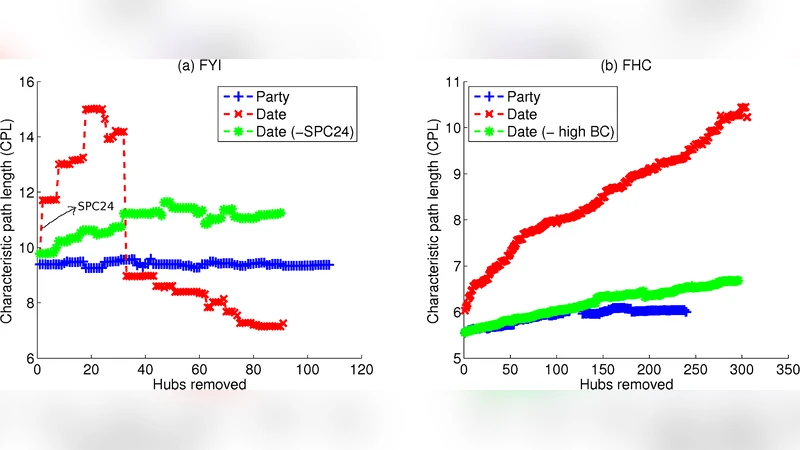
How to understand the cell by breaking it: network analysis of gene perturbation screens

A 3-approximation algorithm for computing a parsimonious first speciation in the gene duplication model
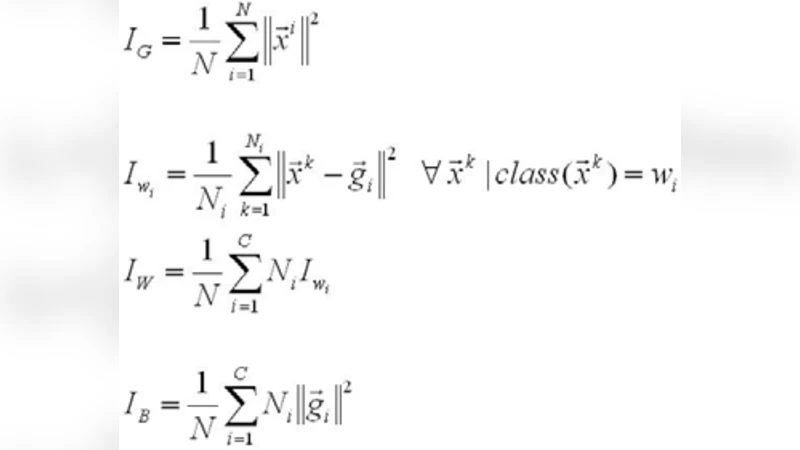
Is it Possible to Extract Metabolic Pathway Information from In Vivo H Nuclear Magnetic Resonance Spectroscopy Data?

Automated Acanthamoeba polyphaga detection and computation of Salmonella typhimurium concentration in spatio-temporal images
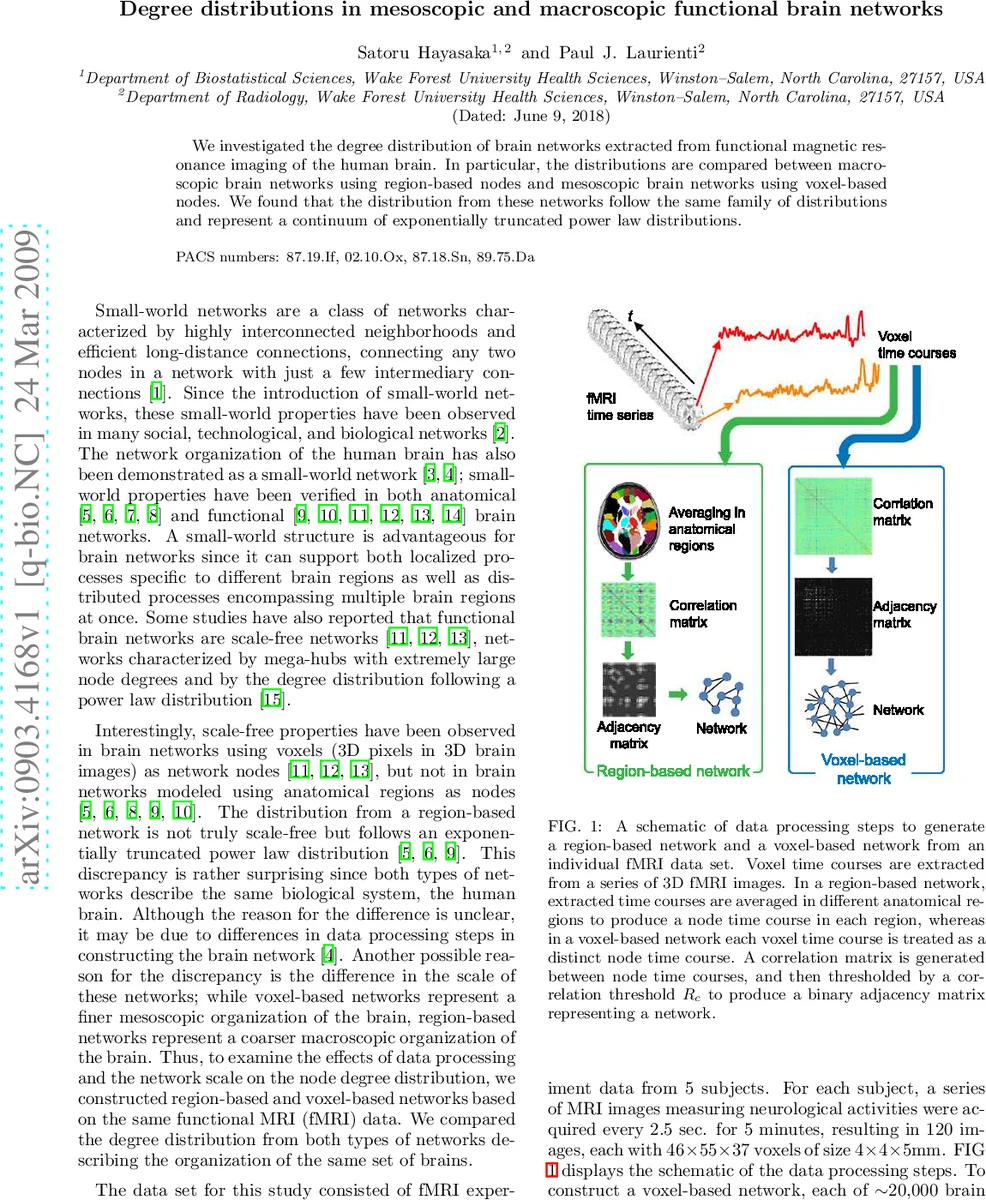

Degree distributions in mesoscopic and macroscopic functional brain networks

Converting genetic network oscillations into somite spatial pattern

Modeling polymerization of microtubules: a quantum mechanical approach

Inherent flexibility determines the transition mechanisms of the EF-hands of Calmodulin

A simple model for dynamic phase transitions in cell spreading

DNA looping provides stability and robustness to the bacteriophage lambda switch

Large attractors in cooperative bi-quadratic Boolean networks. Part I
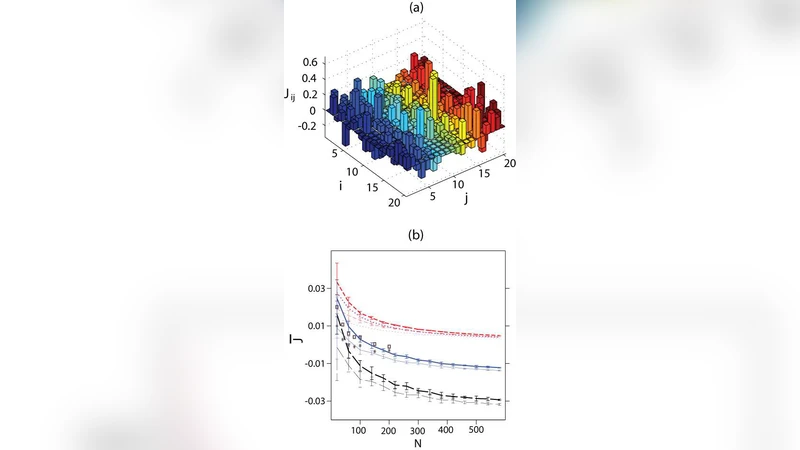
The Ising Model for Neural Data: Model Quality and Approximate Methods for Extracting Functional Connectivity

Comparative analysis of module-based versus direct methods for reverse-engineering transcriptional regulatory networks
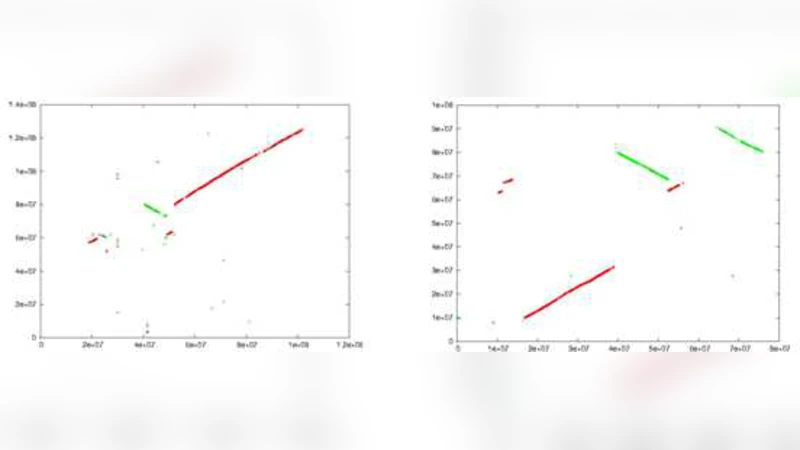
WinBioinfTools: Bioinformatics Tools for Windows High Performance Computing Server 2008

Quantitative test of the barrier nucleosome model for statistical positioning of nucleosomes up- and downstream of transcription start sites

Differential and graphical approaches to multistability theory for chemical reaction networks
