
Q-Bio-Mn

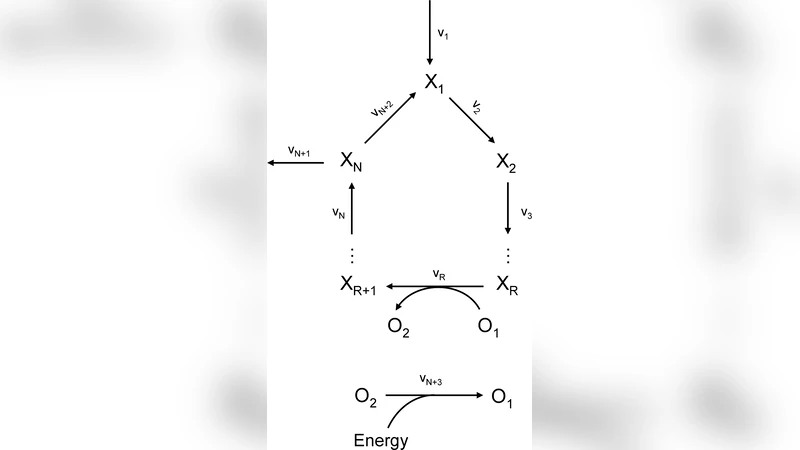
On the Stability of Metabolic Cycles
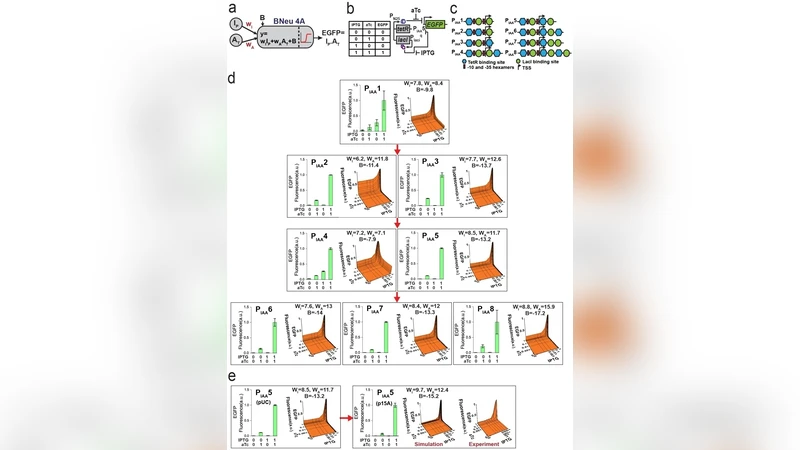
A single layer artificial neural network with engineered bacteria

Binary threshold networks as a natural null model for biological networks
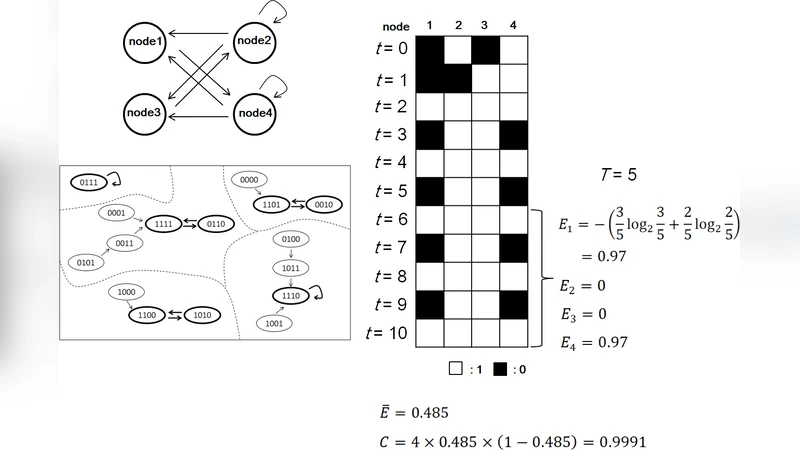
A Multilayer Structure Facilitates the Production of Antifragile Systems in Boolean Network Models

Extremal Properties of Complex Networks

Randomizing genome-scale metabolic networks
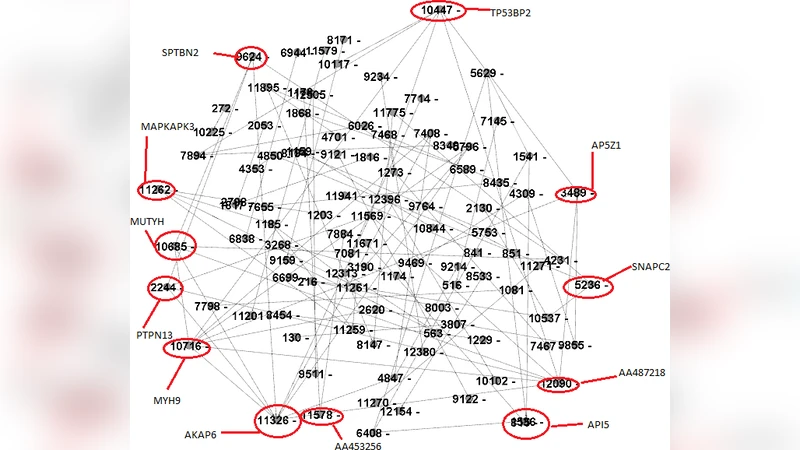
Prediction of a Gene Regulatory Network from Gene Expression Profiles With Linear Regression and Pearson Correlation Coefficient
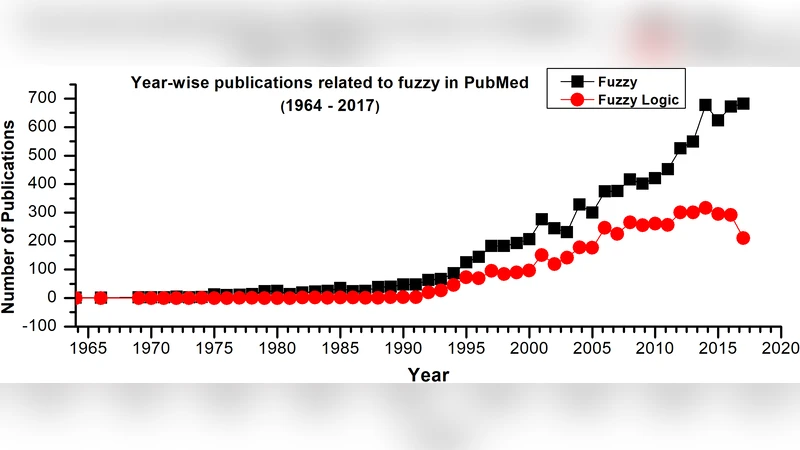
Fuzzy logic based approaches for gene regulatory network inference

Dissecting the Specificity of Protein-Protein Interaction in Bacterial Two-Component Signaling: Orphans and Crosstalks
Estimating Reaction Rate Parameters in Cell Signaling Pathways Using Extreme Decomposition and Belief Propagation Tailored for Data-Rich Cases
Realization and Properties of Biochemical-Computing Biocatalytic XOR Gate Based on Enzyme Inhibition by a Substrate
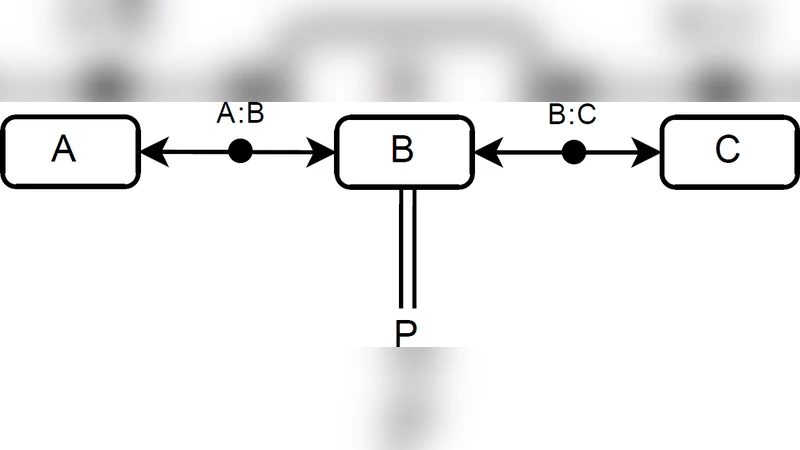
A Process Calculus for Molecular Interaction Maps

Learning of signaling networks: molecular mechanisms

Exploring dimethylsulfate for in vivo proteome footprinting

The Quality of Oscillations in Overdamped Networks

Bayesian design of synthetic biological systems

Optimizing periodicity and polymodality in noise-induced genetic oscillators

Pathways of Distinction Analysis: a new technique for multi-SNP analysis of GWAS data

Cell differentiation: what have we learned in 50 years?
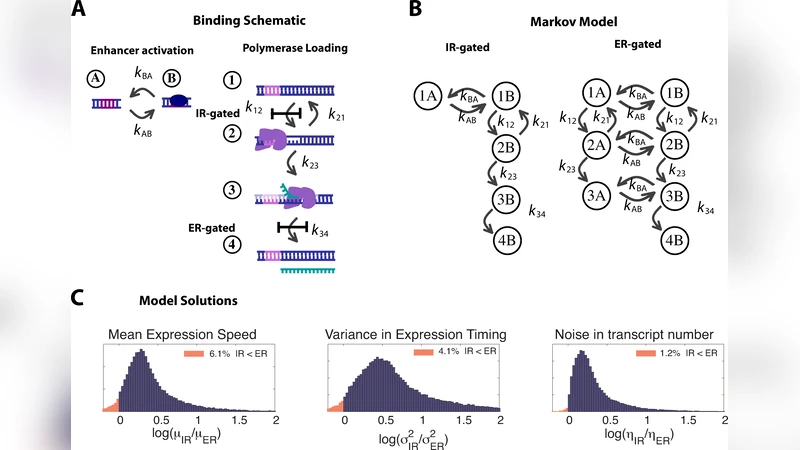
Transcriptional regulation: Effects of promoter proximal pausing on speed, synchrony and reliability

Accurate prediction of gene expression by integration of DNA sequence statistics with detailed modeling of transcription regulation

Motifs emerge from function in model gene regulatory networks
