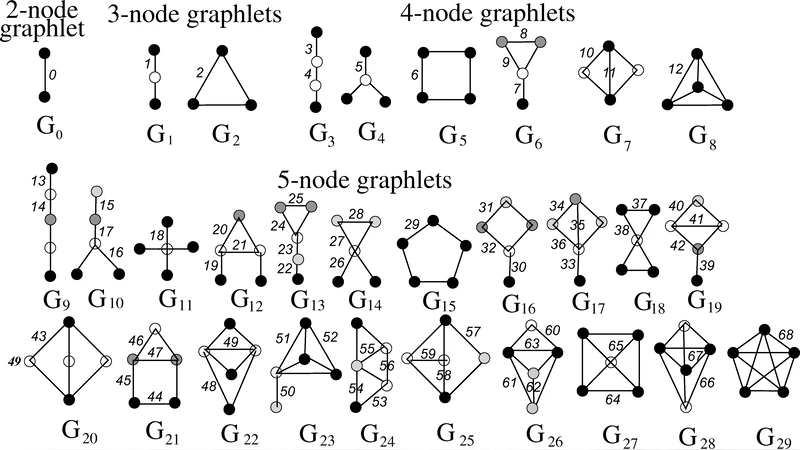
Q-Bio-Mn

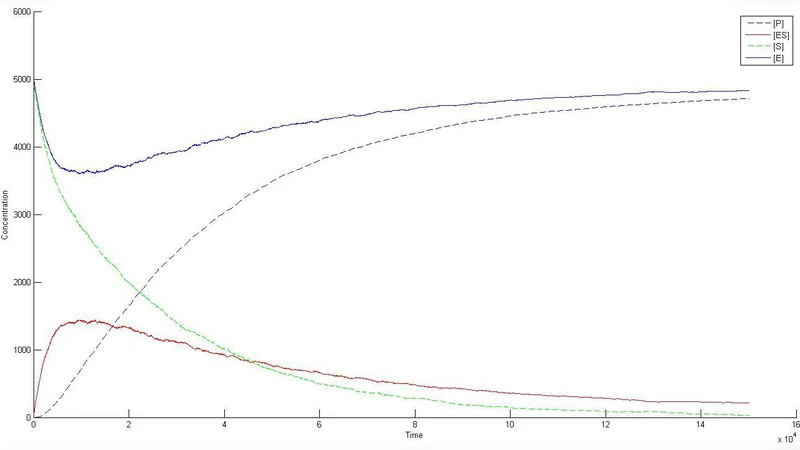
On the Optimization of non-Dense Metabolic Networks in non-Equilibrium State Utilizing 2D-Lattice Simulation

Mining the modular structure of protein interaction networks

Attribute Exploration of Gene Regulatory Processes
The Landscape of Complex Networks

Flexible and robust networks
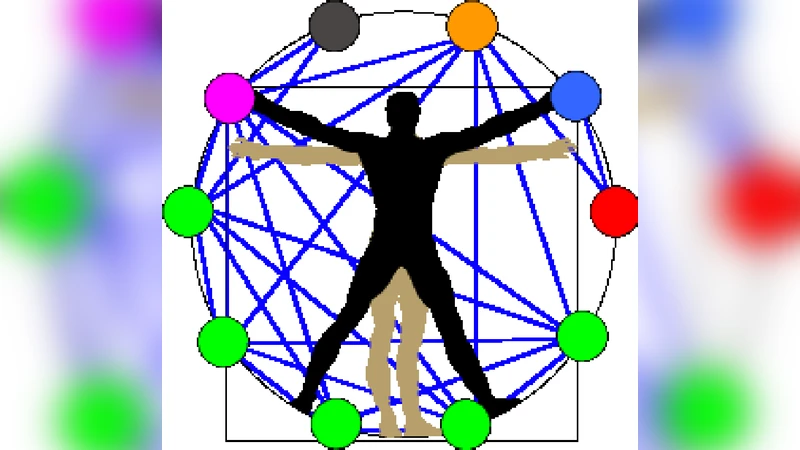
Network Physiology reveals relations between network topology and physiological function

Biochemical pathways simulation
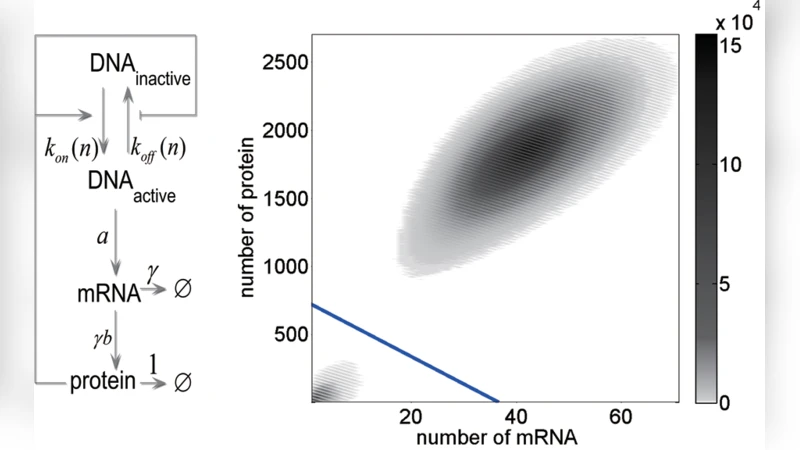
Transition Path, Quasi-potential Energy Landscape and Stability of Genetic Switches
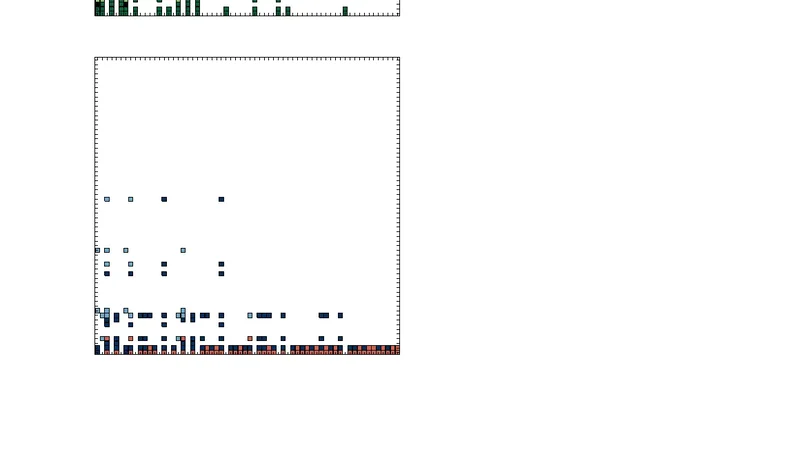
Boolean network-based model of the Bcl-2 family mediated MOMP regulation

Theoretical knock-outs on biological networks
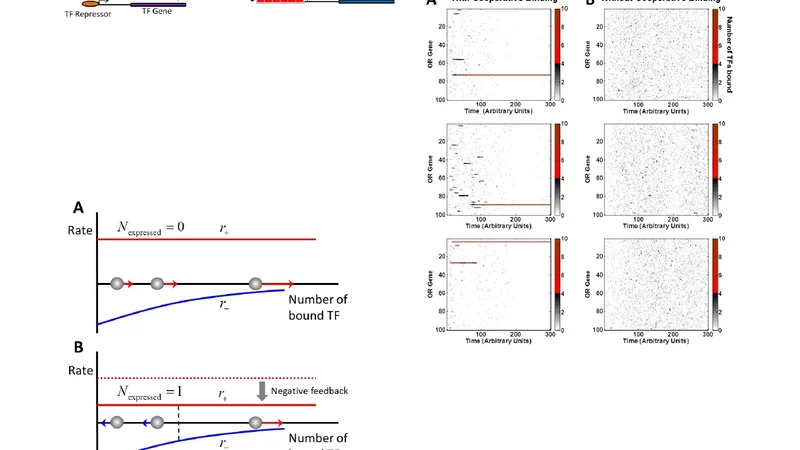
A race model for singular olfactory receptor expression

A stochastic model of B cell affinity maturation and a network model of immune memory

A survey of computational methods for protein complex prediction from protein interaction networks

Control of human immune response function by T-cell population fluctuation and relaxation dynamics
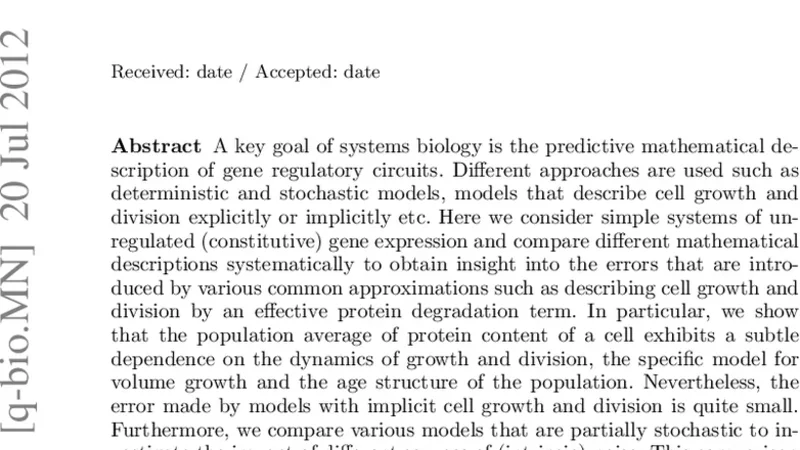

Deterministic and stochastic descriptions of gene expression dynamics

Evolutionary and topological properties of gene modules and driver mutations in a leukemia gene regulatory network
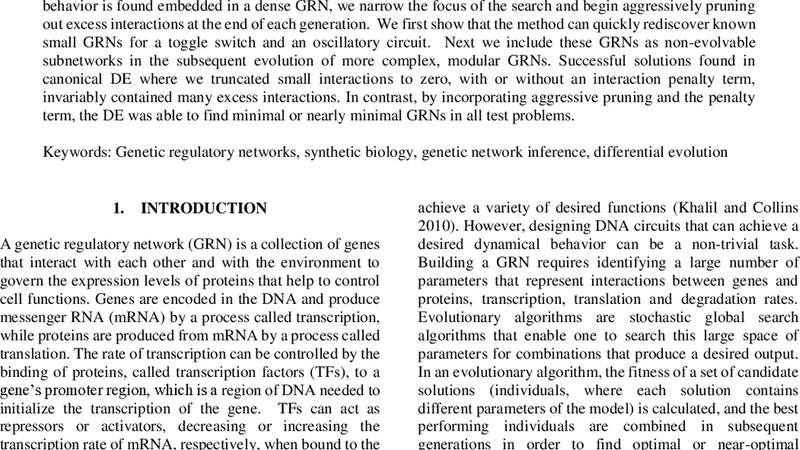

Evolving Modular Genetic Regulatory Networks with a Recursive, Top-Down Approach

Expression proteomics reveals protein targets and highlights mechanisms of action of small molecule drugs
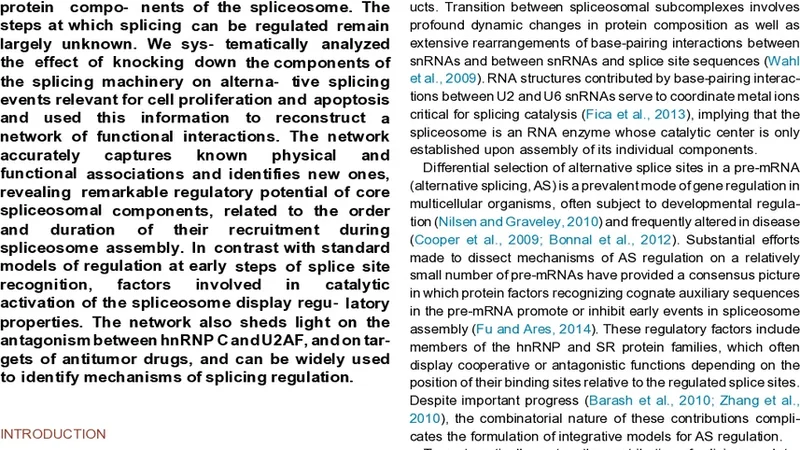

Functional Splicing Network Reveals Extensive Regulatory Potential of the Core Spliceosomal Machinery
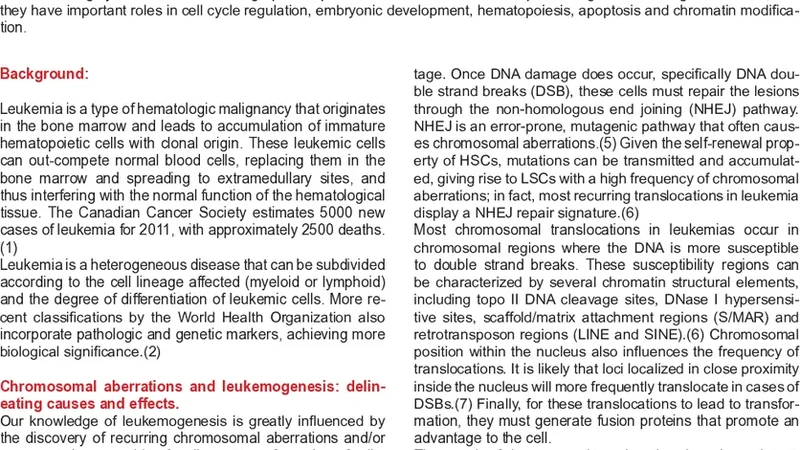

Hematopoietic cancers and Nup98 fusions: determining common mechanisms of malignancy

Identifying robust communities and multi-community nodes by combining top-down and bottom-up approaches to clustering

JAK/STAT signalling - an executable model assembled from molecule-centred modules demonstrating a module-oriented database concept for systems- and synthetic biology
