
Q-Bio-Mn


Optimal Design of a Molecular Recognizer: Molecular Recognition as a Bayesian Signal Detection Problem

Sensitivity analysis of a computational model of the IKK-NF-{kappa}B-I{kappa}B{alpha}-A20 signal transduction network

Controllability, Multiplexing, and Transfer Learning in Networks using Evolutionary Learning

Link between allosteric signal transduction and functional dynamics in a multi-subunit enzyme: S-adenosylhomocysteine hydrolase
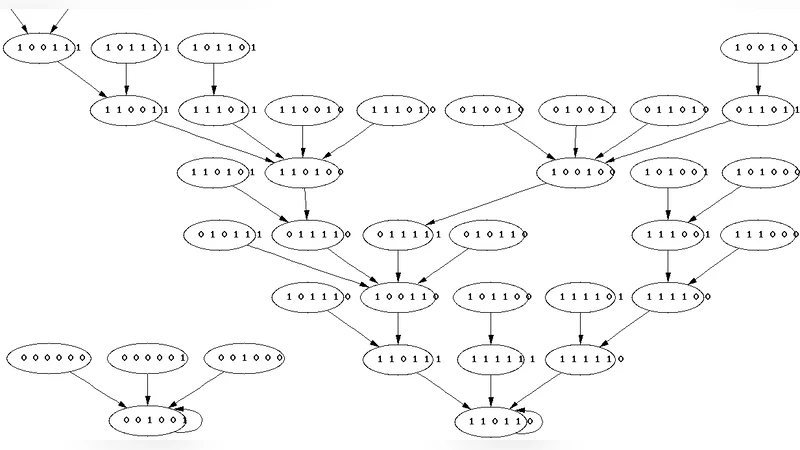
Boolean Models of Bistable Biological Systems
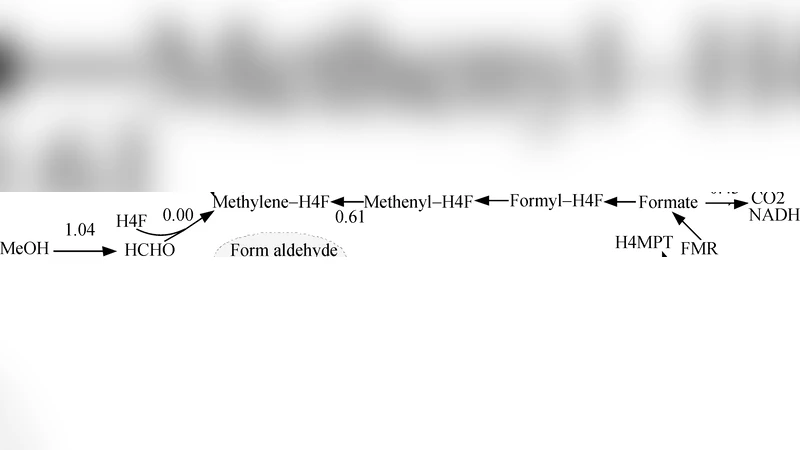
Towards Kinetic Modeling of Global Metabolic Networks with Incomplete Experimental Input on Kinetic Parameters

Specific low frequency electromagnetic fields induce epigenetic and functional changes in U937 cells
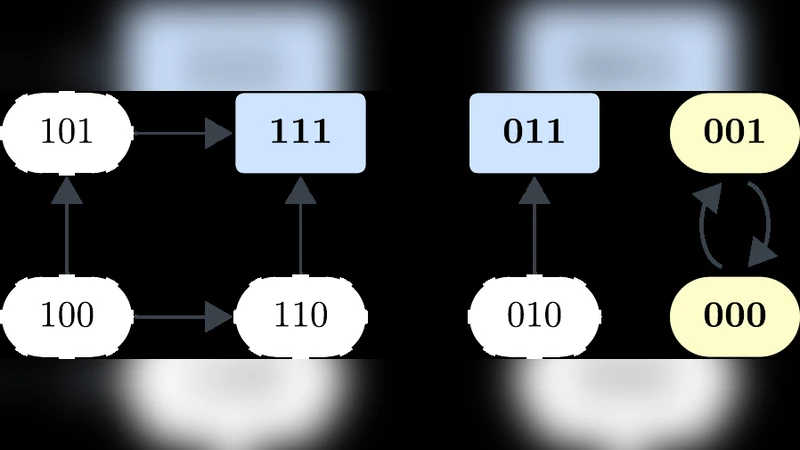
Relating biomarkers and phenotypes using dynamical trap spaces
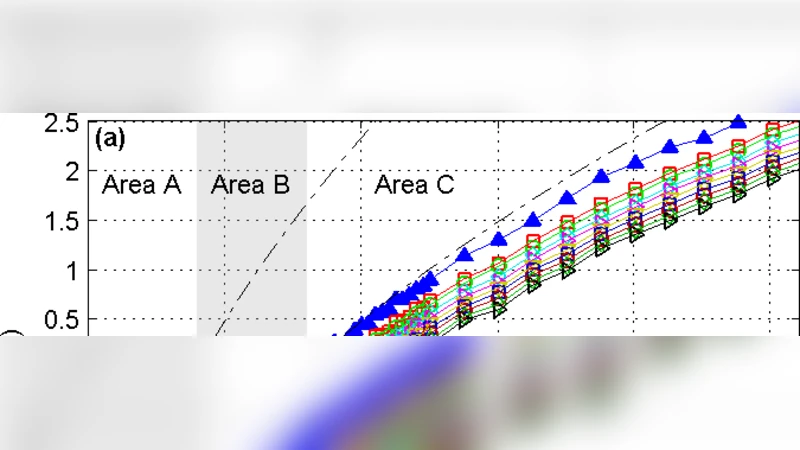
Living on the edge of chaos: minimally nonlinear models of genetic regulatory dynamics

A computational systems biology study of the lambda-lac mutants

Optimizing information flow in small genetic networks. I

Stochastic Control of Metabolic Pathways

Patterns of subnet usage reveal distinct scales of regulation in the transcriptional regulatory network of Escherichia coli
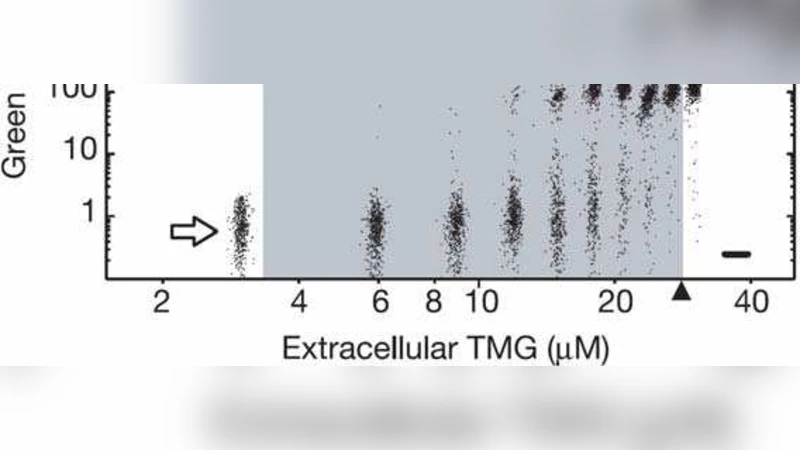
Network Topology as a Driver of Bistability in the lac Operon
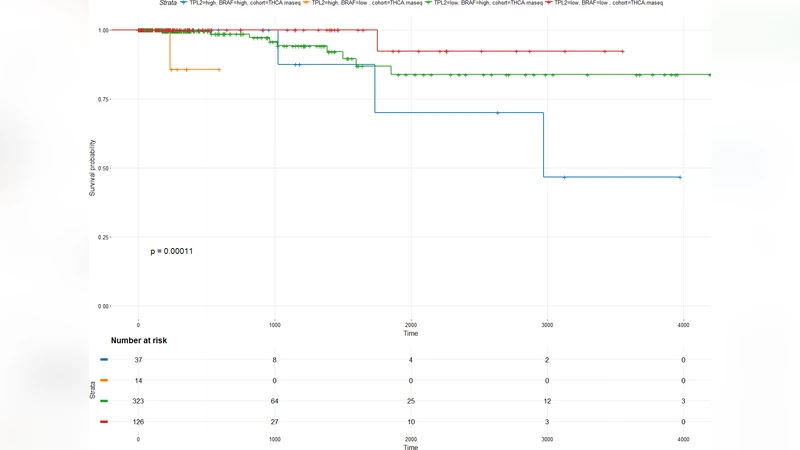
Gene expression and pathway bioinformatics analysis detect a potential predictive value of MAP3K8 in thyroid cancer progression

Flux networks in metabolic graphs

Canalization in the Critical States of Highly Connected Networks of Competing Boolean Nodes

Modular Random Boolean Networks

Evolving Boolean Regulatory Networks with Variable Gene Expression Times

Parameter estimation for Boolean models of biological networks

Predicting disease-related genes by path-based similarity and community structure in protein-protein interaction network

Glutamate regulation of calcium and IP3 oscillating and pulsating dynamics in astrocytes
