Q-Bio-Mn

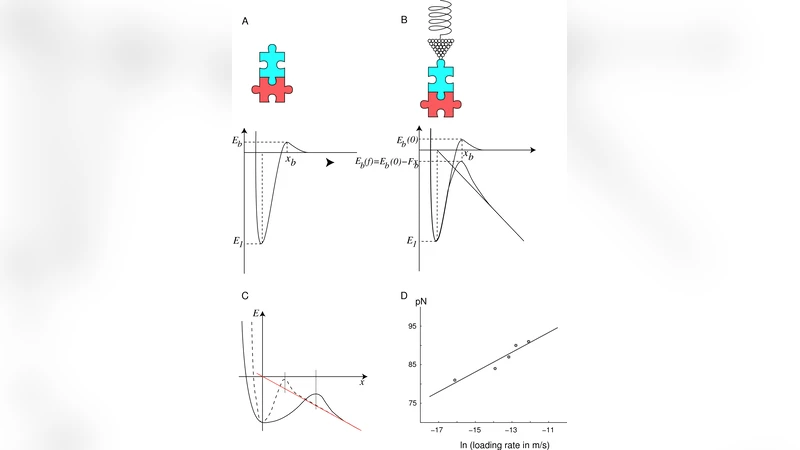
The Interplay between Chemistry and Mechanics in the Transduction of a Mechanical Signal into a Biochemical Function

Quantum information processing at the cellular level. Euclidean approach
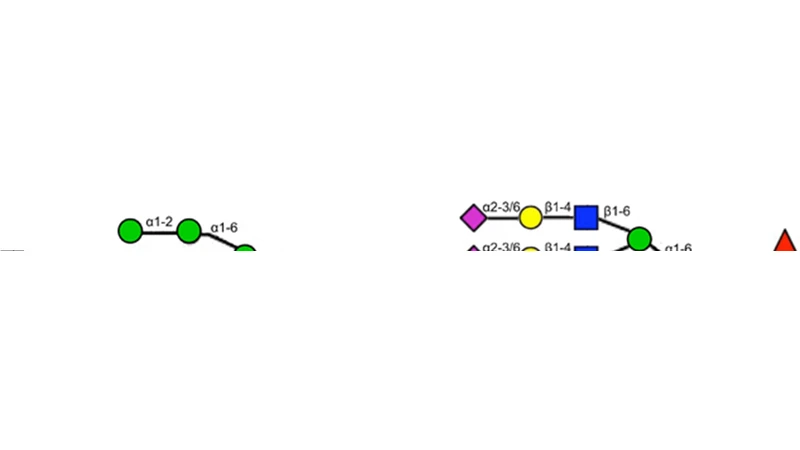
Centralized Modularity of N-Linked Glycosylation Pathways in Mammalian Cells

Global entrainment of transcriptional systems to periodic inputs

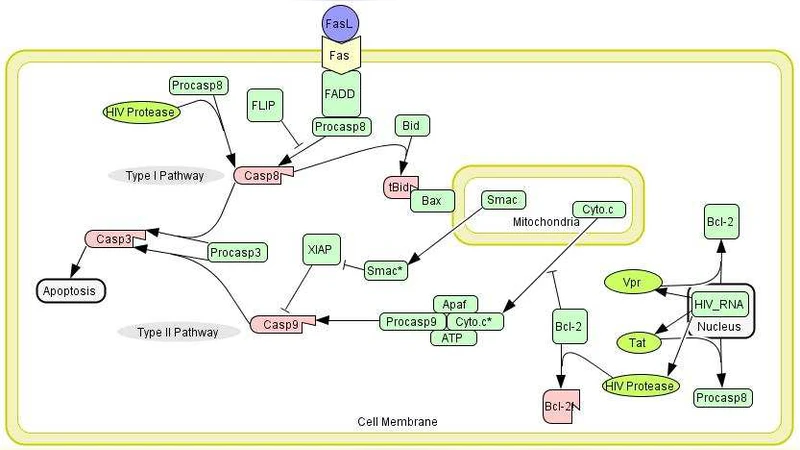
Boolean Network Approach to Negative Feedback Loops of the p53 Pathways: Synchronized Dynamics and Stochastic Limit Cycles

Structure Learning in Nested Effects Models

Partition Decoupling for Multi-gene Analysis of Gene Expression Profiling Data

Reply to Comment on Regularizing Capacity of Metabolic Networks

Equilibrium Blocking Model of Isometric Tension

Allovalency revisited: an analysis of multisite phosphorylation and substrate rebinding

Wide-Scale Analysis of Human Functional Transcription Factor Binding Reveals a Strong Bias towards the Transcription Start Site
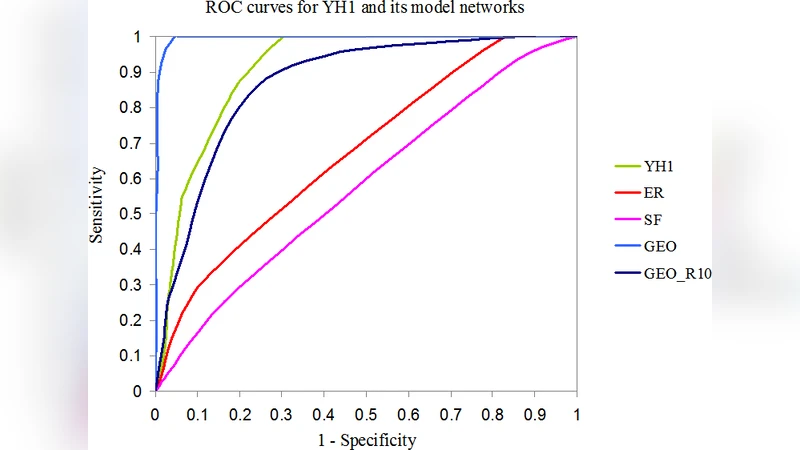
An integrative approach to modeling biological networks

Analytic methods for modeling stochastic regulatory networks

Kinetic Monte Carlo Method for Rule-based Modeling of Biochemical Networks

Network Evolution of Body Plans
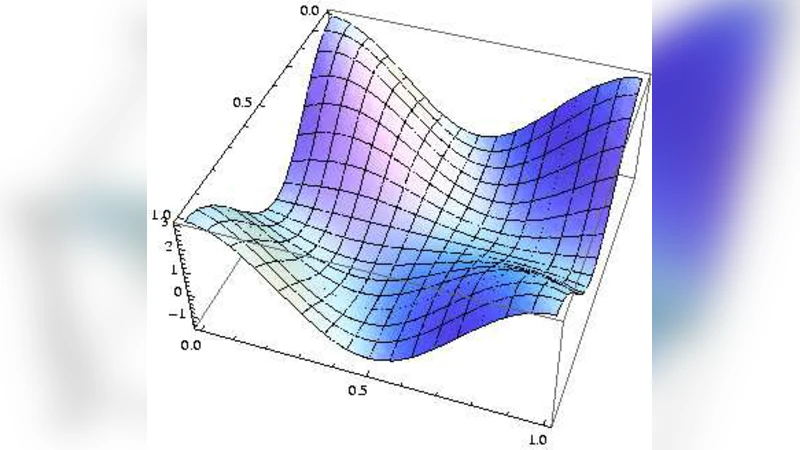
The Repressor-Lattice: Feed-Back, Commensurability, and Dynamical Frustration
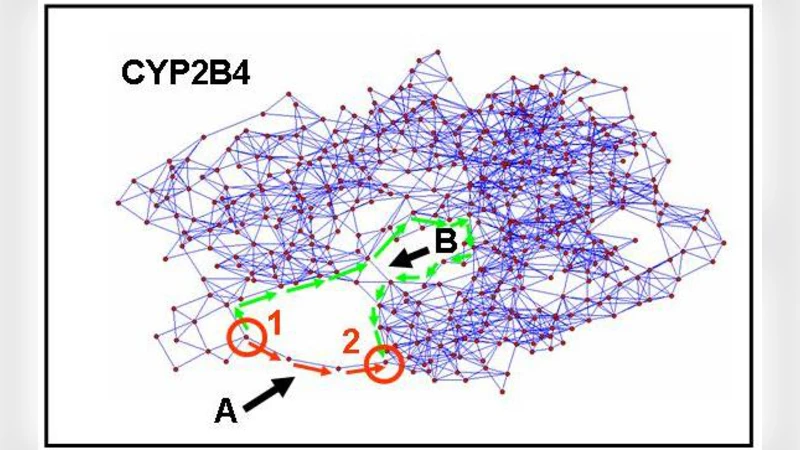
Creative elements: network-based predictions of active centres in proteins, cellular and social networks
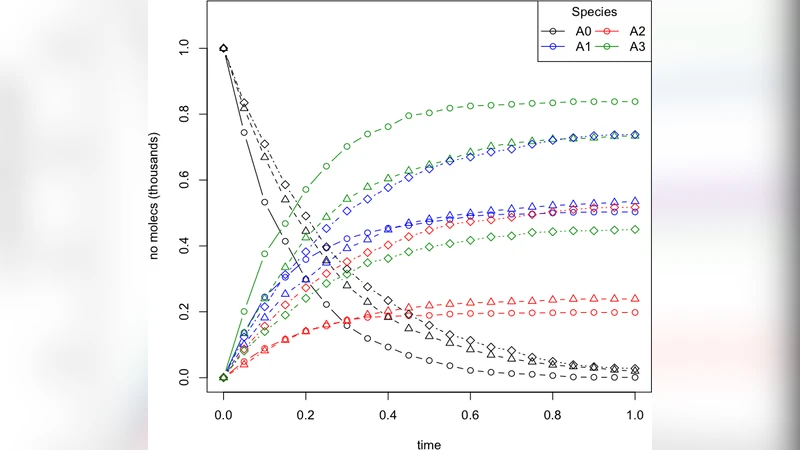
Algebraic Methods for Inferring Biochemical Networks: a Maximum Likelihood Approach

Gene-gene cooperativity in small networks
Toolbox model of evolution of prokaryotic metabolic networks and their regulation

A discrete inhomogeneous model for the yeast cell cycle

A framework for protein and membrane interactions