
Q-Bio-Mn


Functions of Bifans in Context of Multiple Regulatory Motifs in Signaling Networks
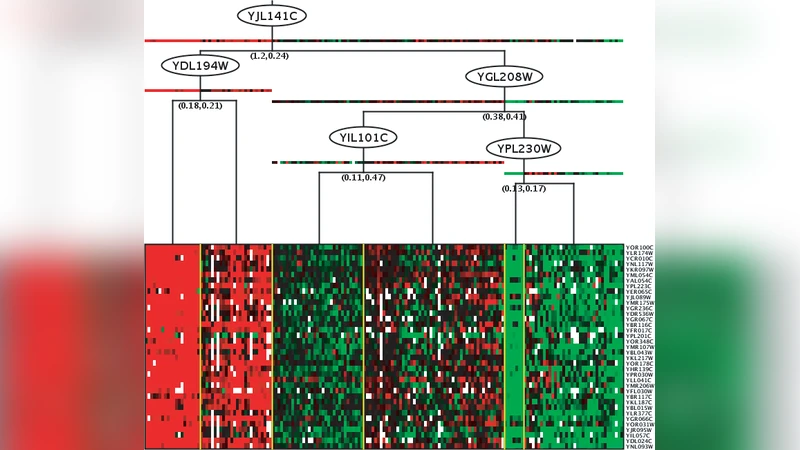
Validating module network learning algorithms using simulated data
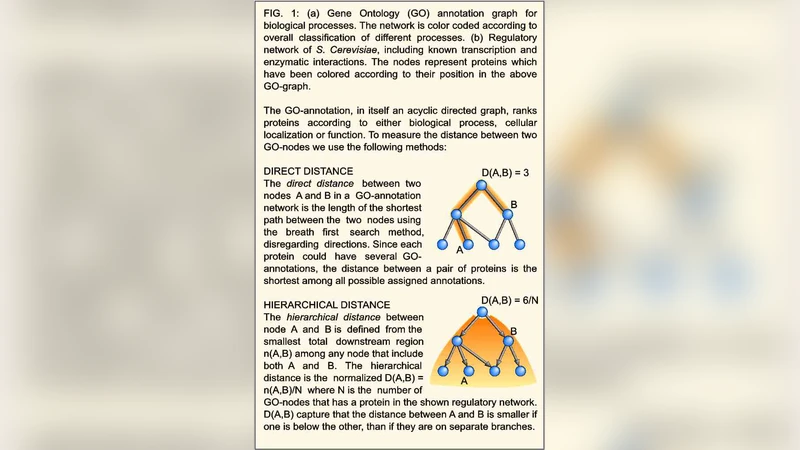

One Hub-One Process: A Tool Based View on Regulatory Network Topology

Epigenetic Chromatin Silencing: Bistability and Front Propagation

Reaction coordinates for the flipping of genetic switches
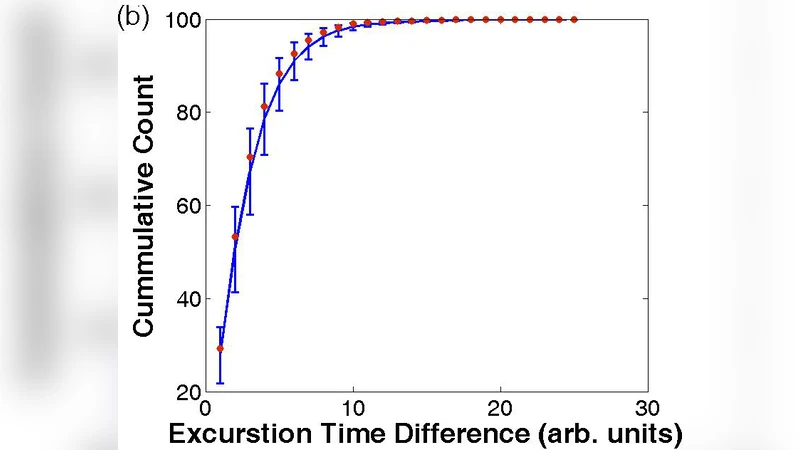
Correlated Phenotypic Transitions to Competence in Bacterial Colonies
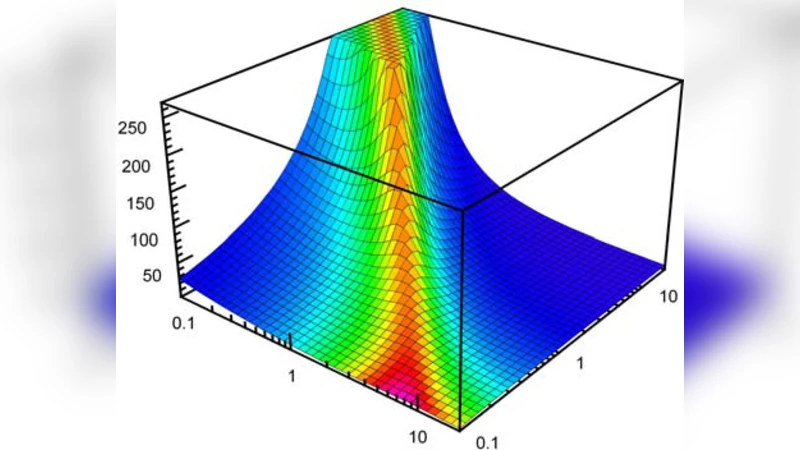
Relationship between cellular response and behavioral variability in bacterial chemotaxis

Information flow and optimization in transcriptional control

Assortative mixing in Protein Contact Networks and protein folding kinetics

Erwin Schroedinger, Francis Crick and epigenetic stability

Network Analysis of Biochemical Logic for Noise Reduction and Stability: A System of Three Coupled Enzymatic AND Gates

Deterministic characterization of stochastic genetic circuits

Colored extrinsic fluctuations and stochastic gene expression

Boolean network model predicts cell cycle sequence of fission yeast

Dynamical Analysis on Gene Activity in the Presence of Repressors and an Interfering Promoter

Multistep greedy algorithm identifies community structure in real-world and computer-generated networks

A quantitative comparison of sRNA-based and protein-based gene regulation
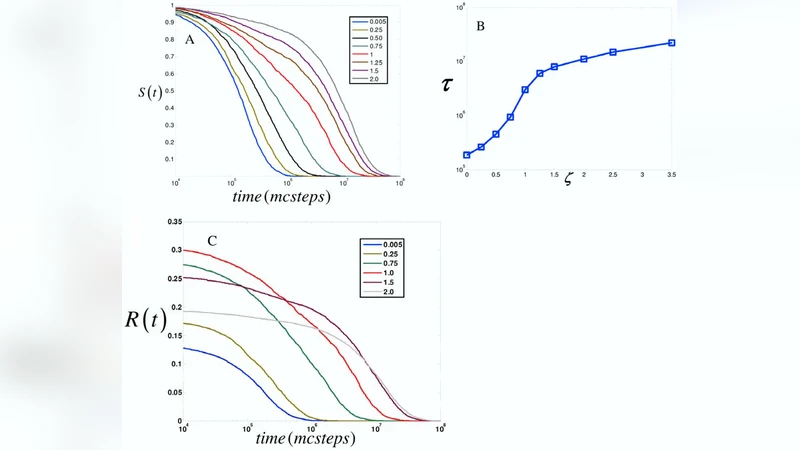
Regulation of signal duration and the statistical dynamics of kinase activation by scaffold proteins
Significance analysis and statistical mechanics: an application to clustering
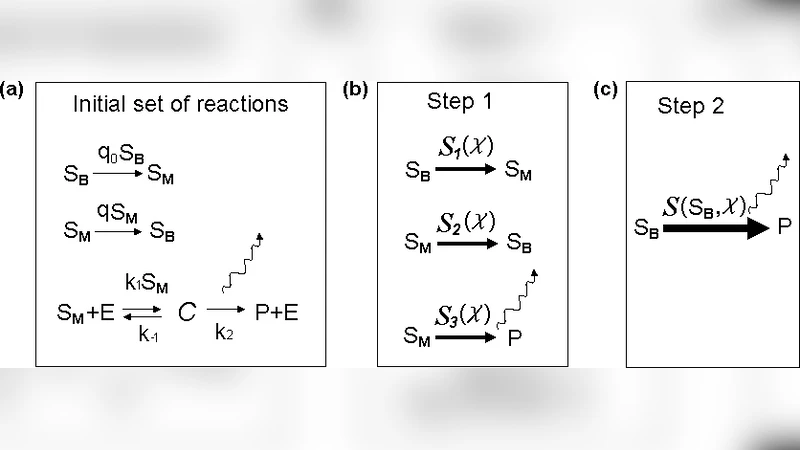
Coarse-graining stochastic biochemical networks: quasi-stationary approximation and fast simulations using a stochastic path integral technique

Effect of promoter architecture on the cell-to-cell variability in gene expression

Protein Interaction Networks are Fragile against Random Attacks and Robust against Malicious Attacks
