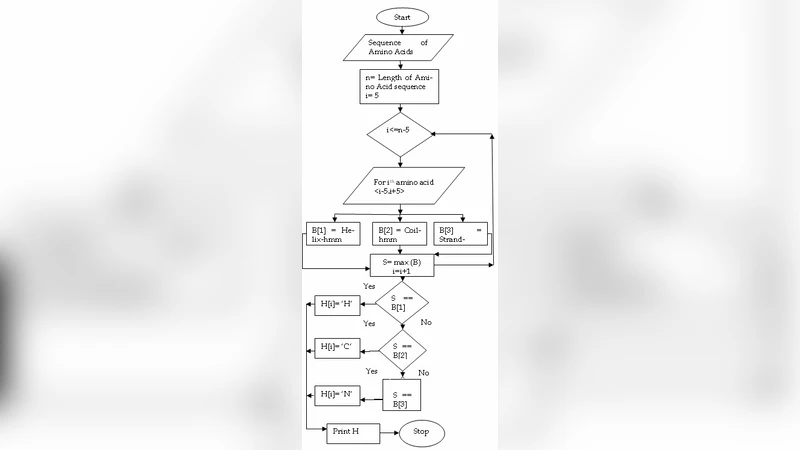
Q-Bio-Bm


Extending fragment-based free energy calculations with library Monte Carlo simulation: Annealing in interaction space
Optical spectrum of proflavine and its ions
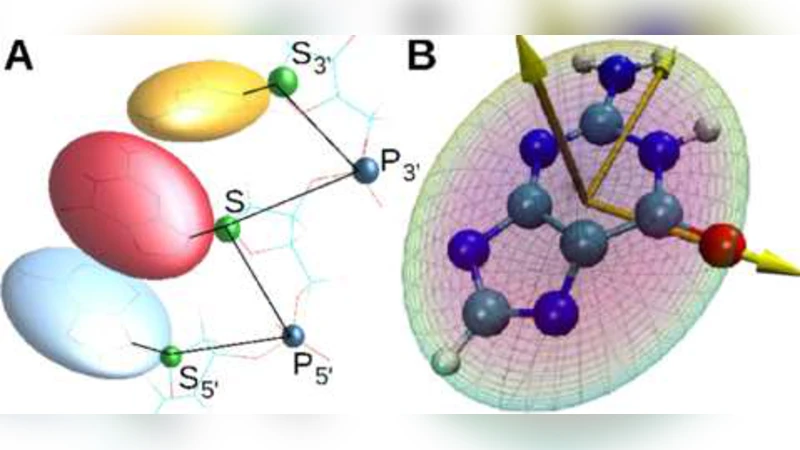
A systematically coarse-grained model for DNA, and its predictions for persistence length, stacking, twist, and chirality

Theoretical Perspectives on Protein Folding

Mathematical models of homochiralisation by grinding of crystals

A simple theory of protein folding kinetics

Inferring DNA sequences from mechanical unzipping data: the large-bandwidth case
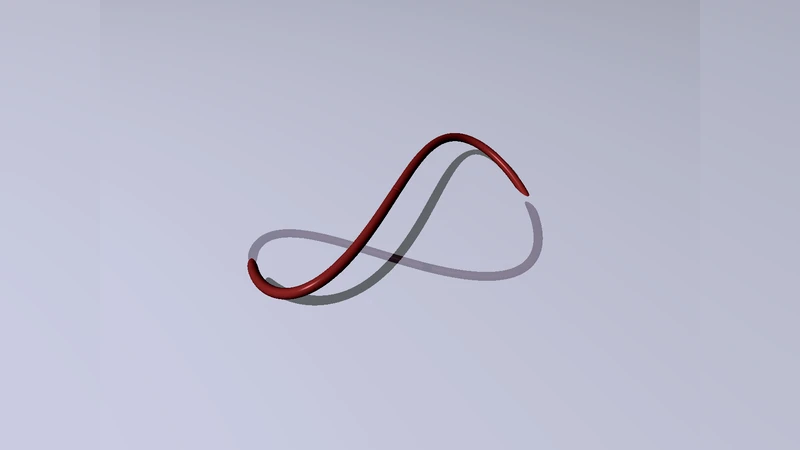

Confinement induces conformational transition of semiflexible polymer rings to figure eight form

Identification of direct residue contacts in protein-protein interaction by message passing
A Finite Element framework for computation of protein normal modes and mechanical response
Self-Templated Nucleation in Peptide and Protein aggregation

Role of ATP-hydrolysis in the dynamics of a single actin filament

Genetic drift at expanding frontiers promotes gene segregation

Contagious obesity: from adenovirus 36 to RB dysfunction

Messenger RNA Fluctuations and Regulatory RNAs Shape the Dynamics of Negative Feedback Loop

The Energy Landscape, Folding Pathways and the Kinetics of a Knotted Protein
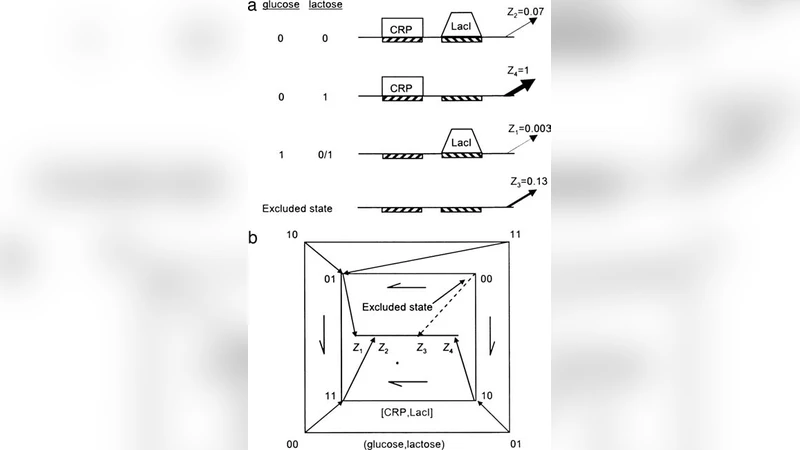
Rules for biological regulation based on error minimization
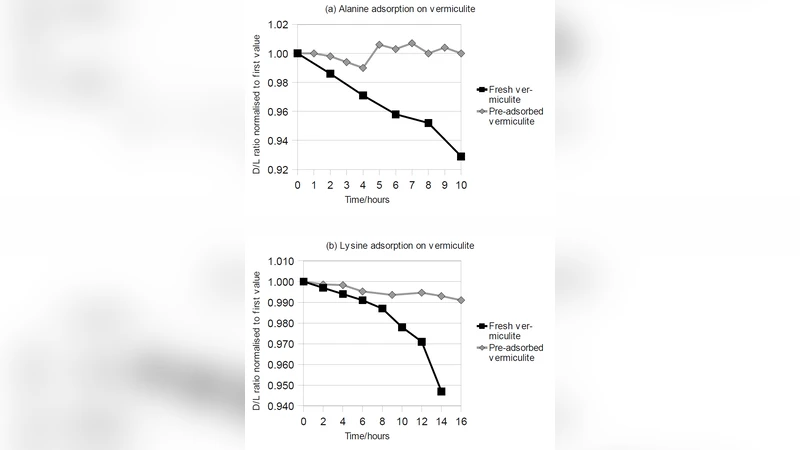
Selective Adsorption and Chiral Amplification of Amino Acids in Vermiculite Clay -Implications for the origin of biochirality

The physical language of molecular codes: A rate-distortion approach to the evolution and emergence of biological codes
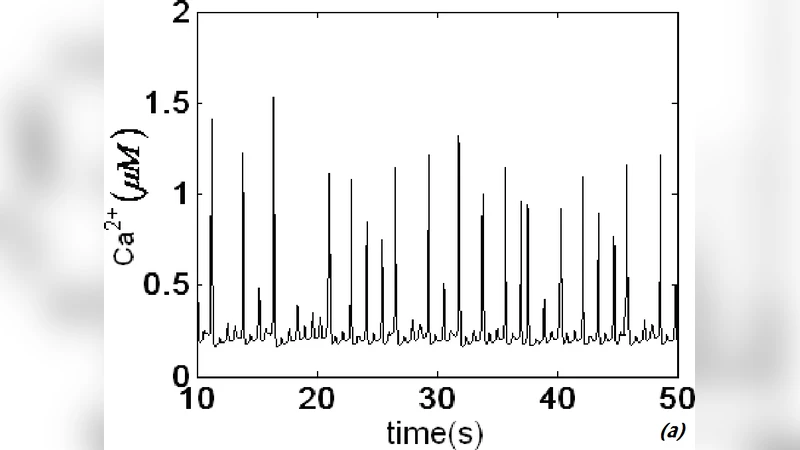
Intercellular spiral waves of calcium in a two dimensional network of cells

Bayesian algorithms for recovering structure from single-particle diffraction snapshots of unknown orientation: a comparison

Probing Noise in Gene Expression and Protein Production
