Q-Bio-Bm


Denaturation Patterns in Heterogeneous DNA

Thermodynamic stability of small-world oscillator networks: A case study of proteins

Optimality Properties of a Proposed Precursor to the Genetic Code

Hidden Semi-Markov Models for Single-Molecule Conformational Dynamics

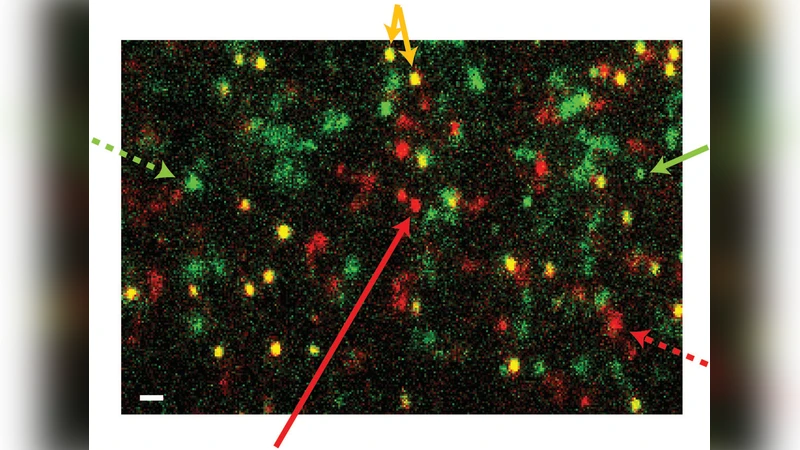
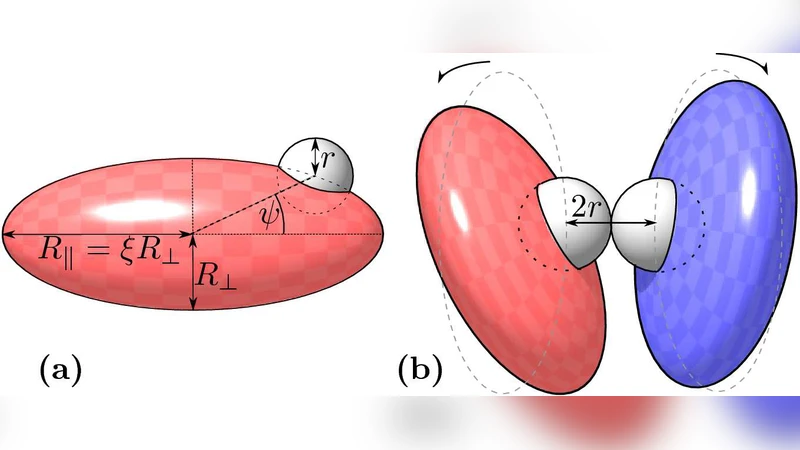
Morphology and Interaction between Lipid Domains
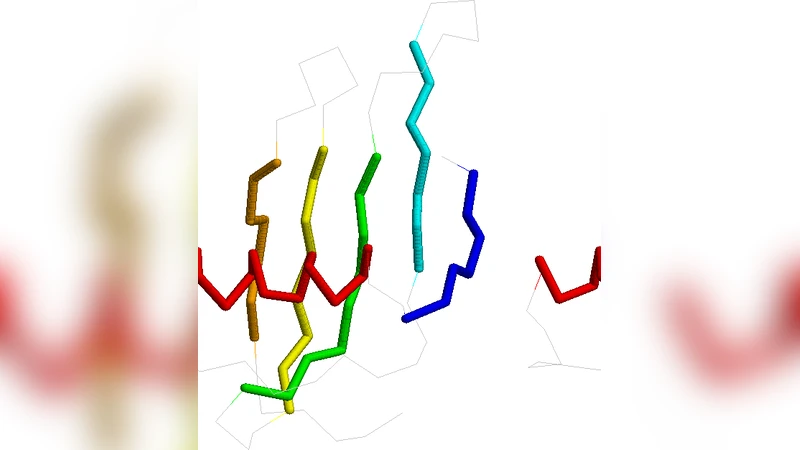
Hydropathy Conformational Letter and its Substitution Matrix HP-CLESUM: an Application to Protein Structural Alignment

Energy landscape of ubiquitin modulated by periodic forces: Asymmetric protein stability and shifts in unfolding pathways
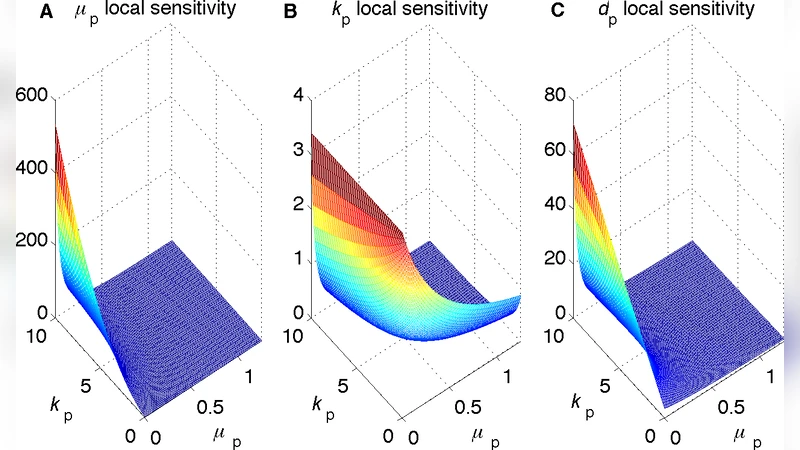
Enhancement of cargo processivity by cooperating molecular motors

Kinetics of the helix-coil transition
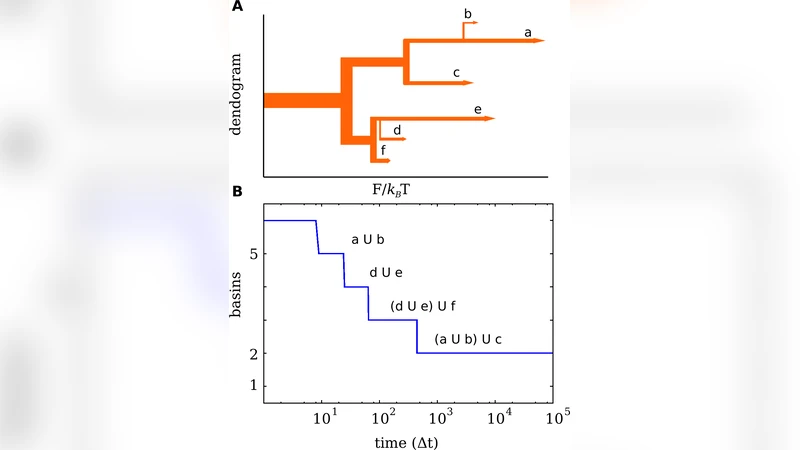
Exploring the Free Energy Landscape: From Dynamics to Networks and Back

Spatial structure and composition of polysaccharide-protein complexes from Small Angle Neutron Scattering
Critical examination of the inherent-structure-landscape analysis of two-state folding proteins
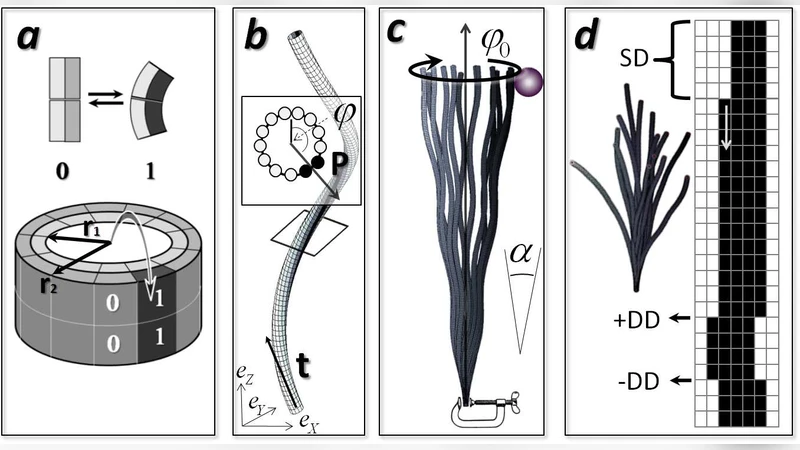
Polymorphic Dynamics of Microtubules

Stochastic resonance of ELF-EMF in voltage-gated channels: the case of the cardiac I_Ks potassium channel
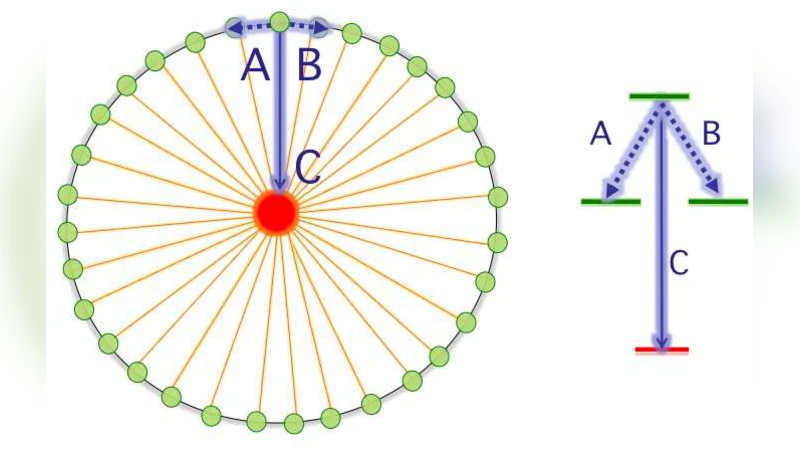
Non-Markovian stochastic description of quantum transport in photosynthetic systems
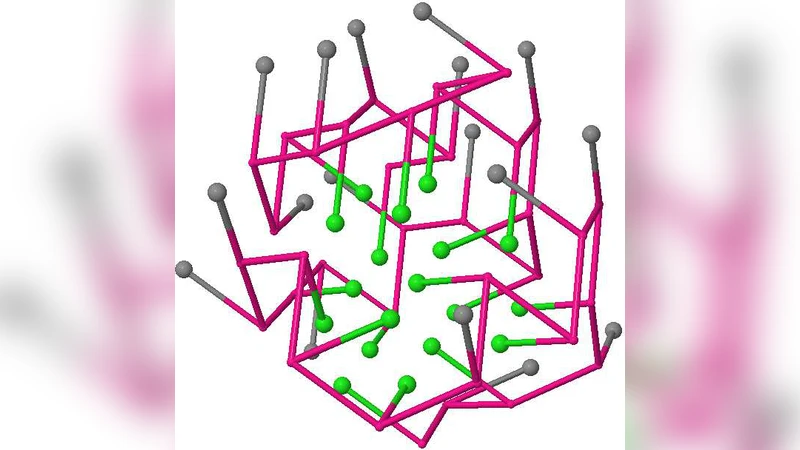
Equivalence Classes of Optimal Structures in HP Protein Models Including Side Chains
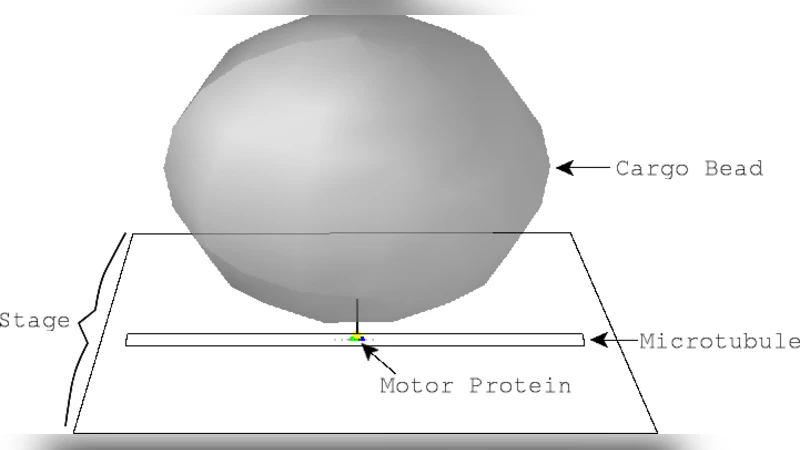
A Brownian Dynamics Model of Kinesin in Three Dimensions Incorporating the Force-Extension Profile of the Coiled-Coil Cargo Tether

Protein Folding: A Perspective From Statistical Physics
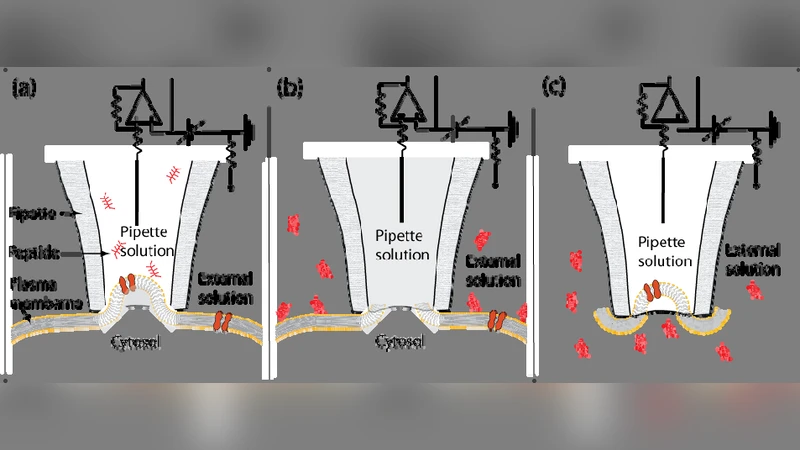
Arginine-rich peptides destabilize the plasma membrane, consistent with a pore formation translocation mechanism of cell penetrating peptides
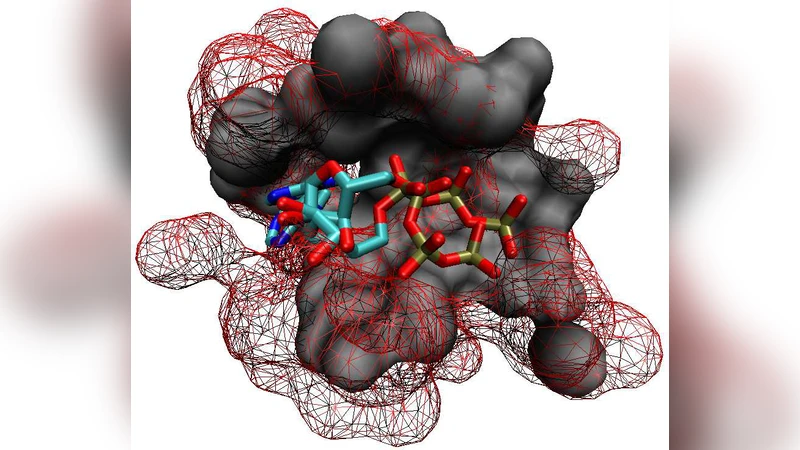
A new protein binding pocket similarity measure based on comparison of 3D atom clouds: application to ligand prediction

Simulating Stochastic Dynamics Using Large Time Steps
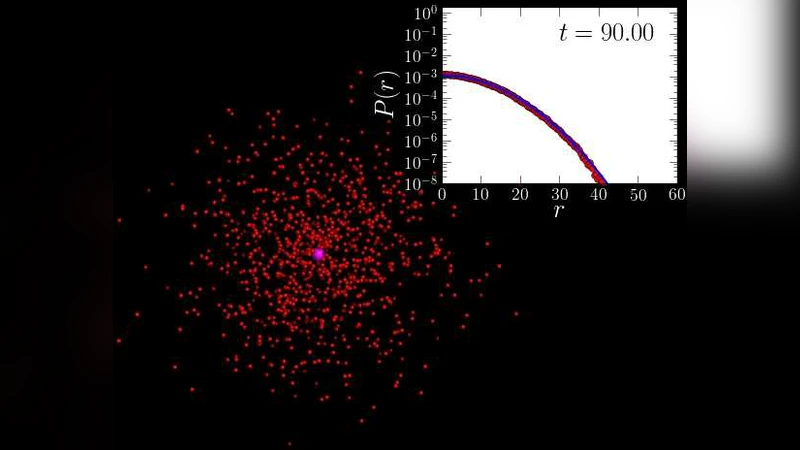
DNA-Protein Binding Rates: Bending Fluctuation and Hydrodynamic Coupling Effects