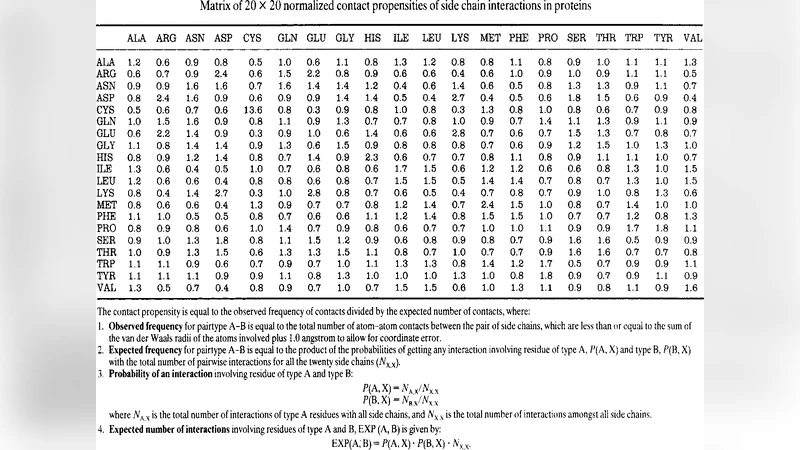
Q-Bio-Bm


Minimal models for proteins and RNA: From folding to function

Bayesian analysis of time series of single RNA under fluctuating force

Two-State Folding, Folding through Intermediates, and Metastability in a Minimalistic Hydrophobic-Polar Model for Proteins
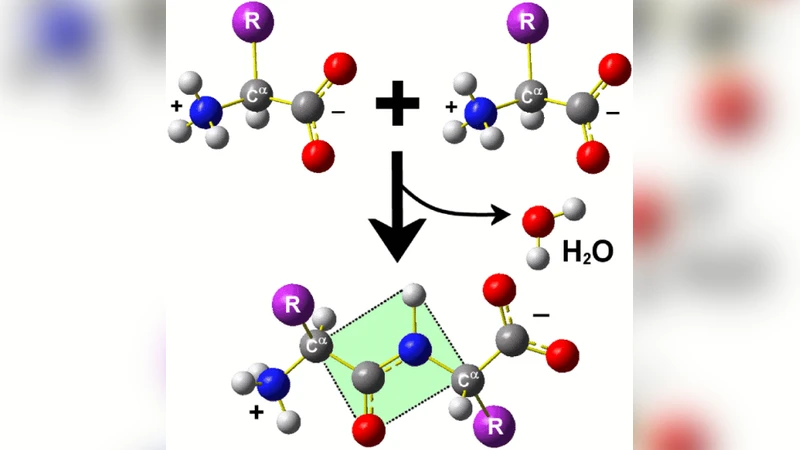
Introduction to protein folding for physicists

Similarity search for local protein structures at atomic resolution by exploiting a database management system

Monte Carlo simulations of proteins in cages: influence of confinement on the stability of intermediate states

Pore-blockade Times for Field-Driven Polymer Translocation

Peeling and Sliding in Nucleosome Repositioning
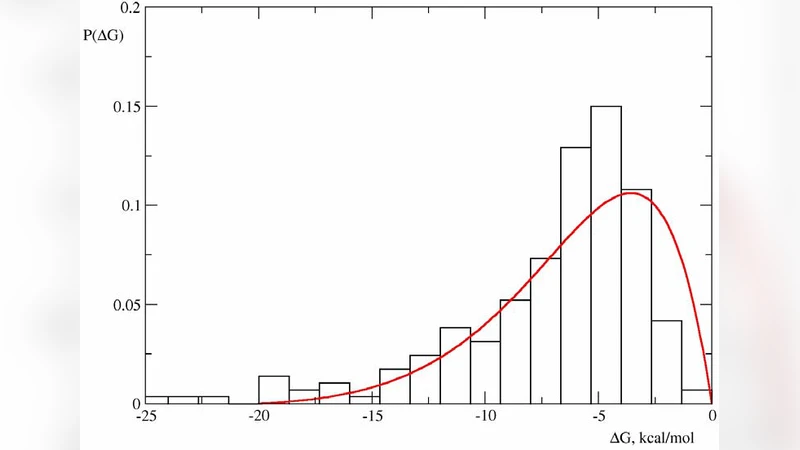
The Hypercube of Life: How Protein Stability Imposes Limits on Organism Complexity and Speed of Molecular Evolution

Metadynamic sampling of the free energy landscapes of proteins coupled with a Monte Carlo algorithm

Assortative mixing in Protein Contact Networks and protein folding kinetics

Reconstructing the free energy landscape of a mechanically unfolded model protein
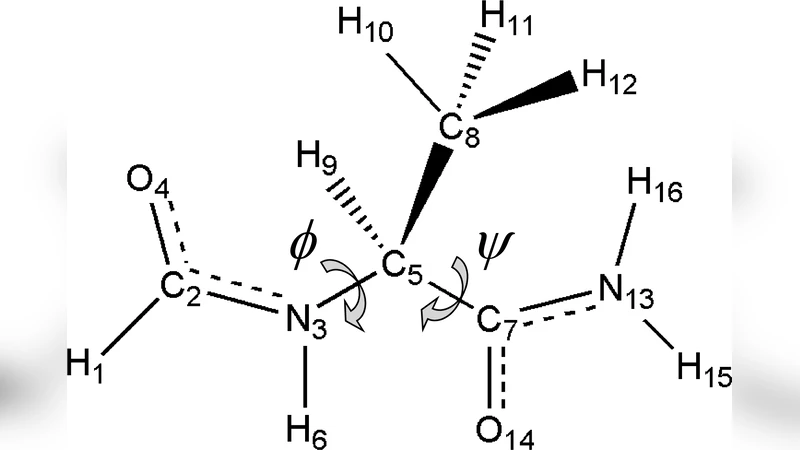
Efficient model chemistries for peptides. I. Split-valence Gaussian basis sets and the heterolevel approximation in RHF and MP2
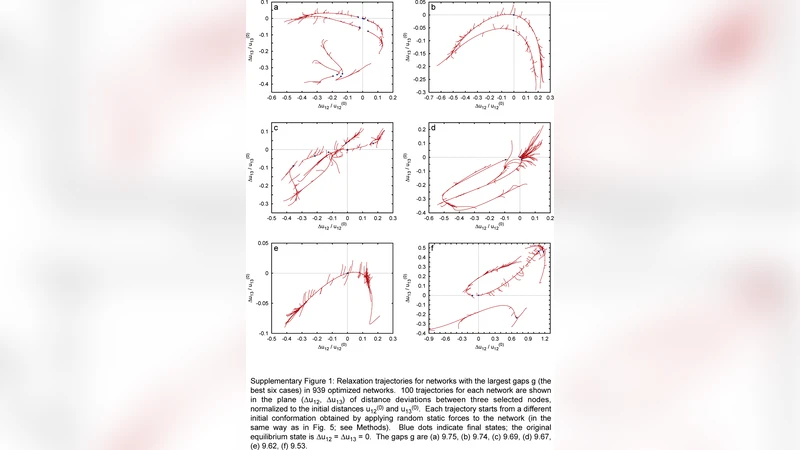
Nonlinear Relaxation Dynamics in Elastic Networks and Design Principles of Molecular Machines

AFM Imaging of SWI/SNF action: mapping the nucleosome remodeling and sliding
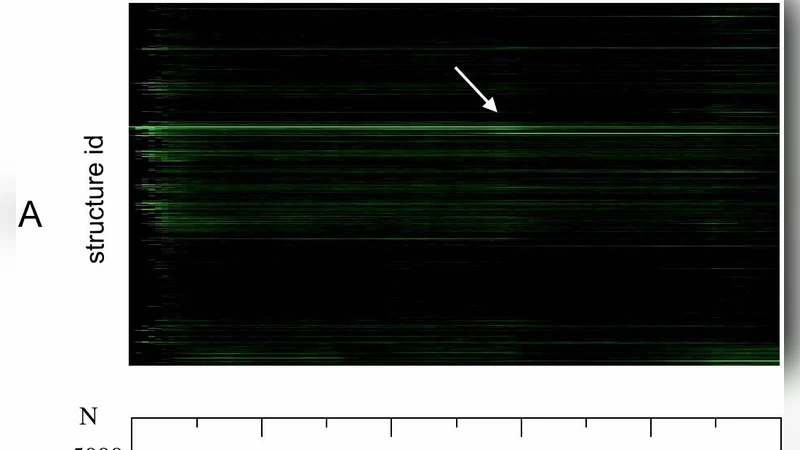
A first-principles model of early evolution: Emergence of gene families, species and preferred protein folds
Coupling of transverse and longitudinal response in stiff polymers
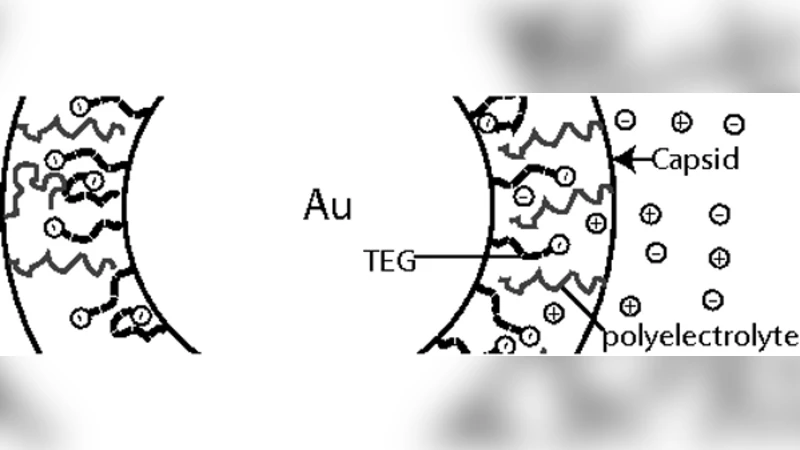
A theory for viral capsid assembly around electrostatic cores

Functioning of the dimeric GABA(B) receptor extracellular domain revealed by glycan wedge scanning

Atomic Biology, Electrostatics, and Ionic Channels

Spontaneous Unknotting of a Polymer Confined in a Nanochannel

Predicting Transcription Factor Specificity with All-Atom Models
