Q-Bio-Bm


Modelling the Self-Assembly of Virus Capsids

Maximum Cliques in Protein Structure Comparison
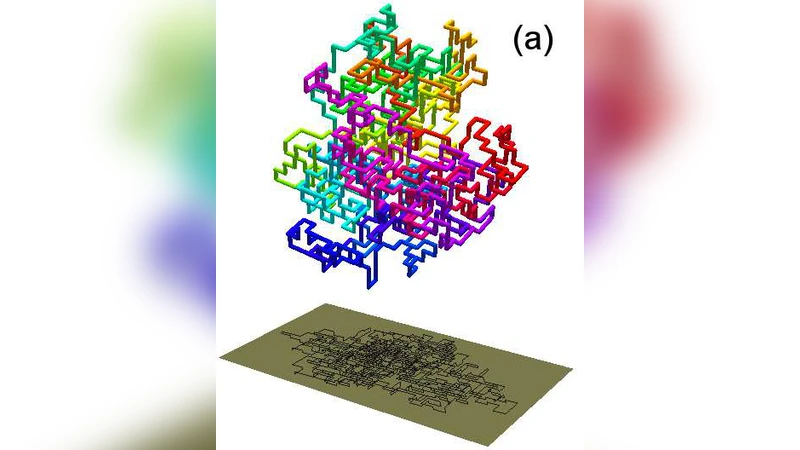
Interplay between writhe and knotting for swollen and compact polymers

Mechanical Strength of 17 134 Model Proteins and Cysteine Slipknots

Mechanical model for a collagen fibril pair in extracellular matrix
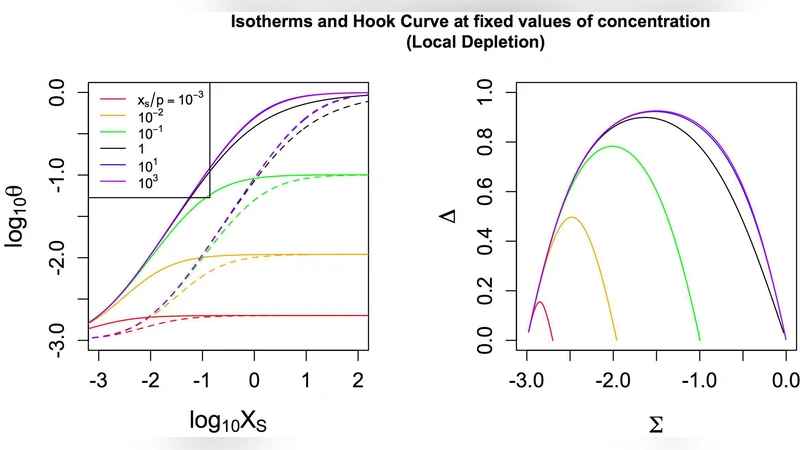
Physico-chemical modelling of target depletion during hybridisation on oligonulceotide microarrays
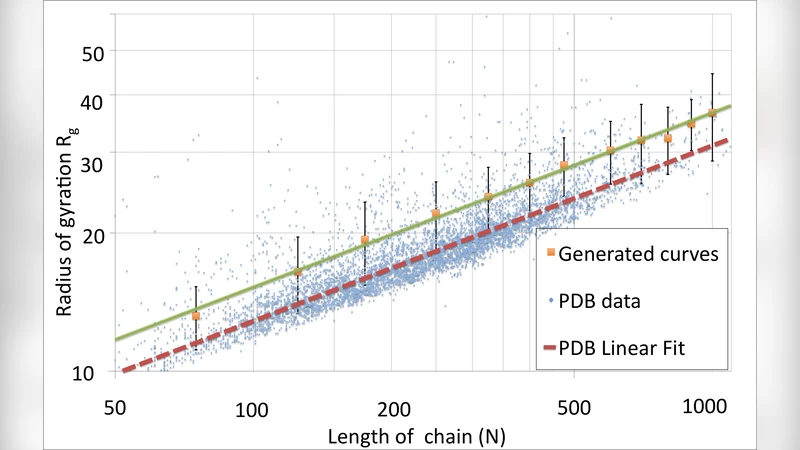
A Gauge Field Theory of Chirally Folded Homopolymers with Applications to Folded Proteins
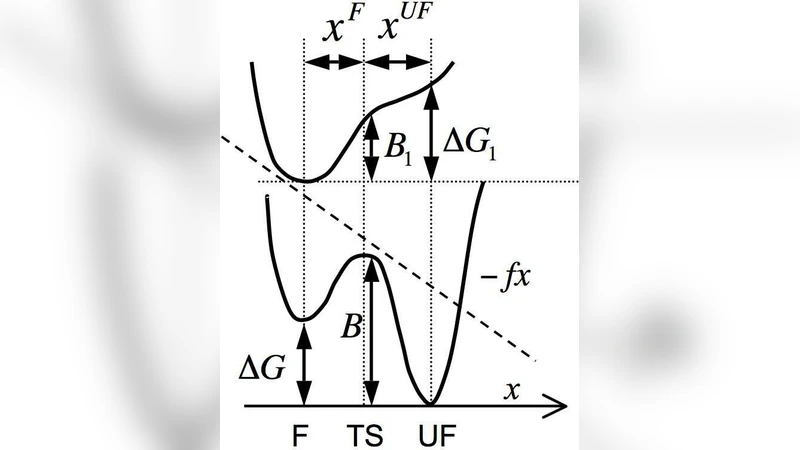
Dynamic force spectroscopy of DNA hairpins. II. Irreversibility and dissipation

Multi-Dimensional Theory of Protein Folding

Influence of Protein Electromagnetic Field on Hydrogen Bonding
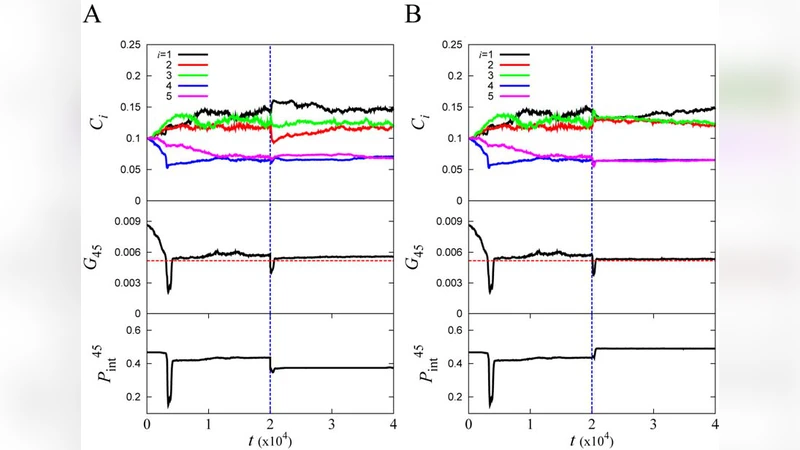
Adaptation through stochastic switching into transient mutators in finite asexual populations

Fatgraph Models of Proteins
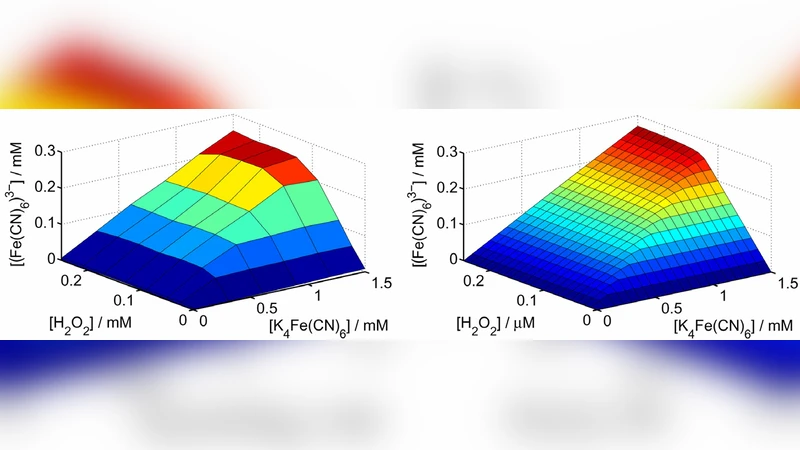
Optimization of Enzymatic Logic Gates and Networks for Noise Reduction and Stability

Excluded volume effects on semiflexible ring polymers

Propagation of twist solitons in real DNA chains

Atomic-detailed milestones along the folding trajectory of protein G

Analog Noise Reduction in Enzymatic Logic Gates
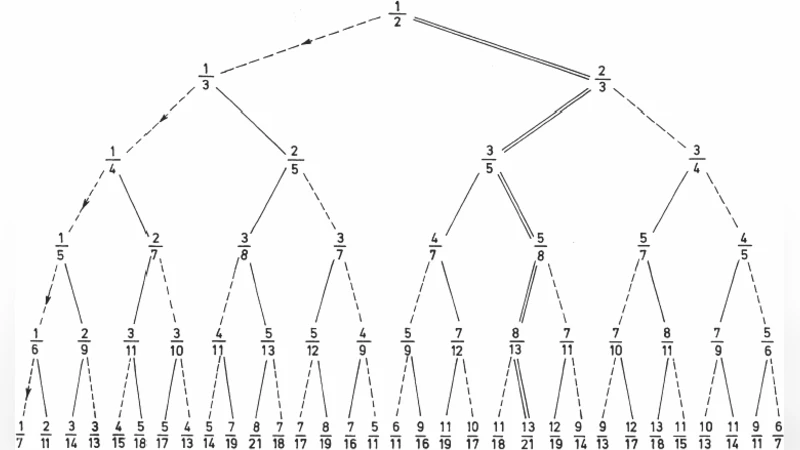
Genetic Code Table: A note on the three splittings into amino acid classes
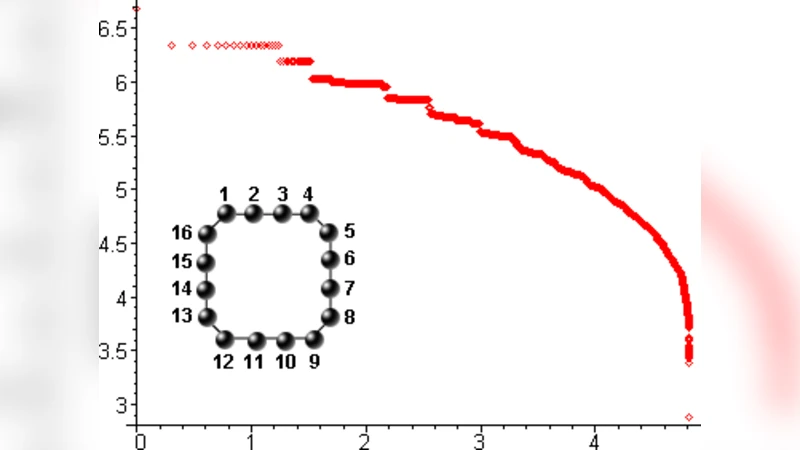
Neutral Networks of Sequence to Shape Maps
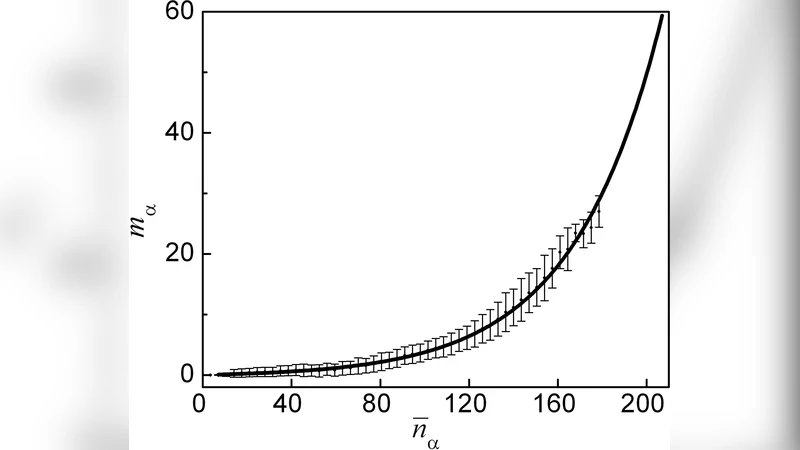
Hidden Structure in Protein Energy Landscapes
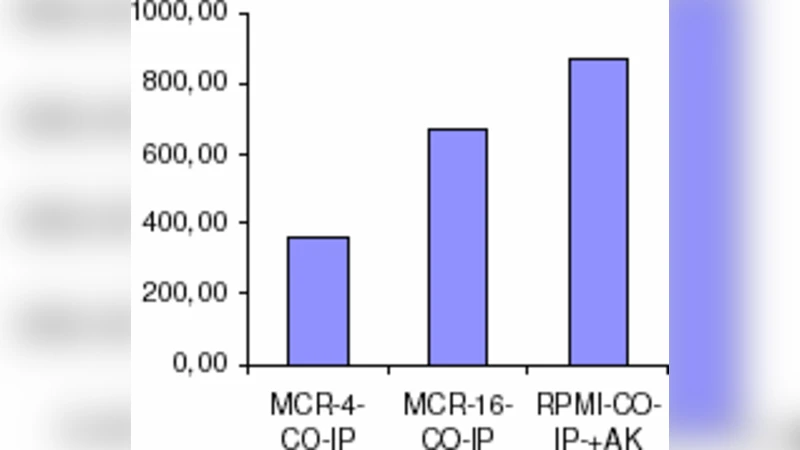
Antiproliferative MCR peptides block physical interaction of insulin with retinoblastoma protein (RB) in human lung cancer cells

Biodegradation of 2-ethylhexyl nitrate (2-EHN) by Mycobacterium austroafricanum IFP 2173
